A patient of his forwarded me the Metagenomics Stool results from Laboratorium voor Medische Ontledingen. I have created a page to derive recommendations from it: Metagenomics Stool (De Meirleir).
Report
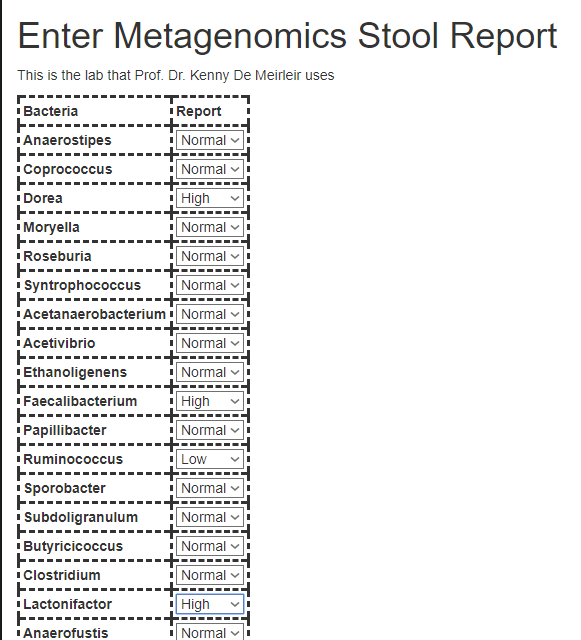
This is a long report with over 50 genus having ranges. An example is below

Enter Page
As with other lab tests, there is a simple entry page to record high, low and normals. The order is the same as on the report.
Recommendations
Note: If nothing appears, go back and try again — odd bug has appeared.
See Making sense of Avoid and Take on Recommendations

Bottom Line
More self-serve for people working with De Meirleir.
This is an education post to facilitate discussing this approach with your medical professionals. It is not medical advice for the treatment of any condition. Always consult with your medical professional before doing any changes of diet, supplements or activity. Some items cites may interfere with prescription medicines.