I got an interesting request
I wonder if you would be willing to write a blog post looking at my recent test results in comparison to last year? Confirmed diagnosis of ME/CFS. UK NHS only helps with pacing advice.
- First BiomeSight test: 2025-04-17
Following this, 3 self-directed cycles of antibiotics, probiotics, prebiotics, and diet changes based on MicrobiomePrescription results. First 2 cycles increased my baseline and reduced symptoms dramatically, third cycle set me back slightly. Overall very positive.
Unfortunately then was hospitalised later in 2025 with a perforated and infected gallbladder, sepsis. They rotated through quite a few different harsh antibiotics trying to find one which worked. Then in December 2025 went in for surgery to remove the gallbladder, more antibiotics.
- Second BiomeSight test: 2026-03-16
My baseline now is worse again, many symptoms returned. I am loathe to use more antibiotics while some of my bacteria are so low (Akkermansia at 0.006) even though my positive scores are dominated by antibiotic suggestions. Would like to focus on probiotics, prebiotics, herbs, supplements, diet changes for now.
Any insight would be most appreciated.
Confirming the Worst
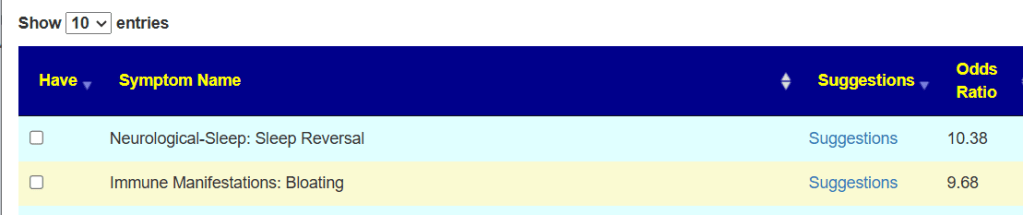
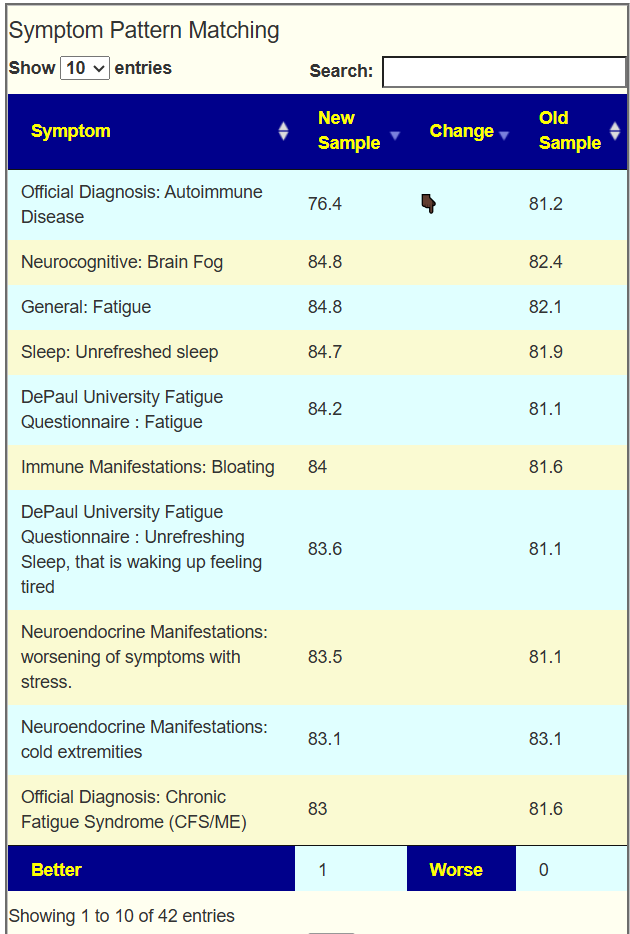
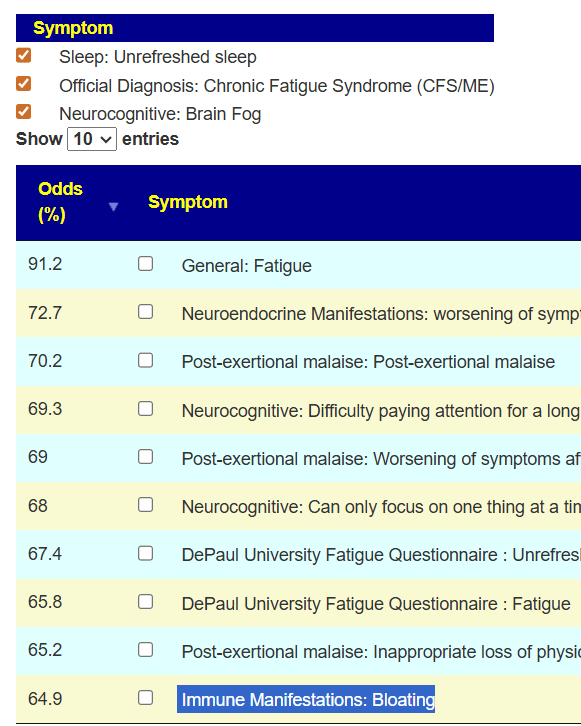
Going over to the symptom compare tool, we see that you are now worse than a year ago. 140 of 141 symptom forecasts are significantly worse! Seeing number this much worse is unusual but consistent with his events and perception.
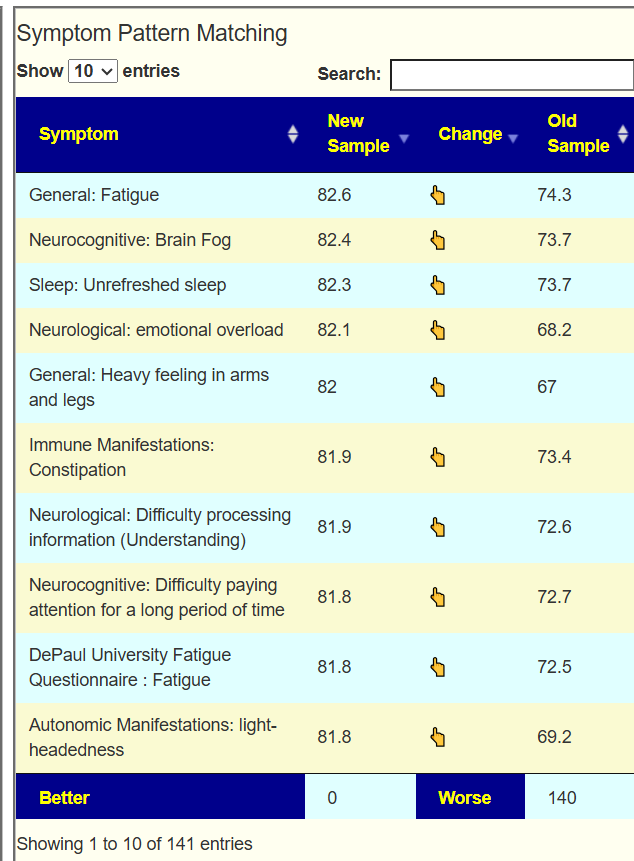
Going Forward
My last two post has been evaluating the alternative path — instead of attacking the bacteria causing symptoms, push the person to a statistically significant healthy microbiome. The following links may be worth a reading:
- Another ME/CFS Microbiome Update – Review of a sample
- A new sample and two roads to walk – Review of a sample
- Good vs Bad Bacteria Dogmatic Beliefs – background
- Turning Fixing the Microbiome Upside Down! – background
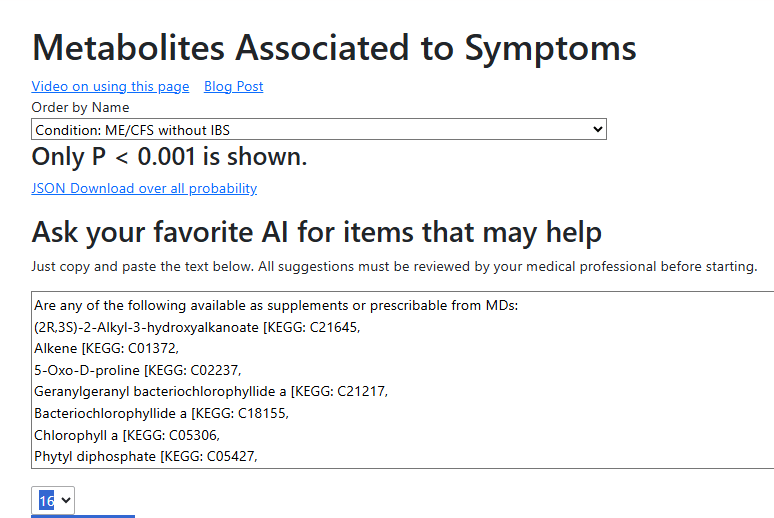
This approach matches “I am loathe to use more antibiotics” because antibiotics typically are on the avoid lists with the healthy approach and high on the to take list attacking symptoms. It is sitting on the Simple UI page.

Basic Results:
- 52 bacteria were identified — every single one was too high.
- I have broken suggestions into classes below. In general, I have kept them to items with an impact of at least 1.
- Items listed are order by largest impact first.
Herbs
The top herbs are below. I was delighted {Bofutsushosan} was listed because it is well known increases Akkermansia which he is concerned about.

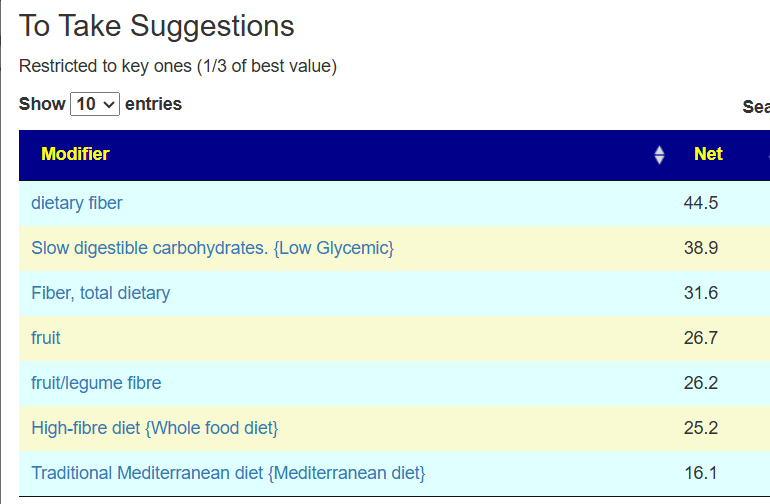
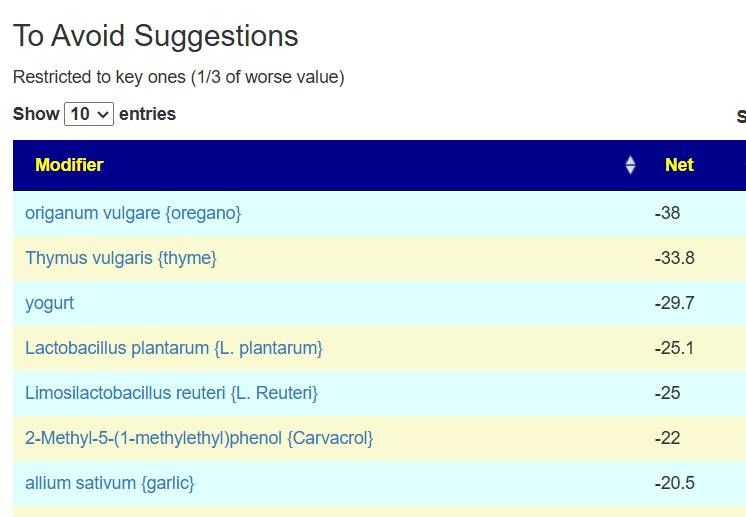
Food

Flavonoids
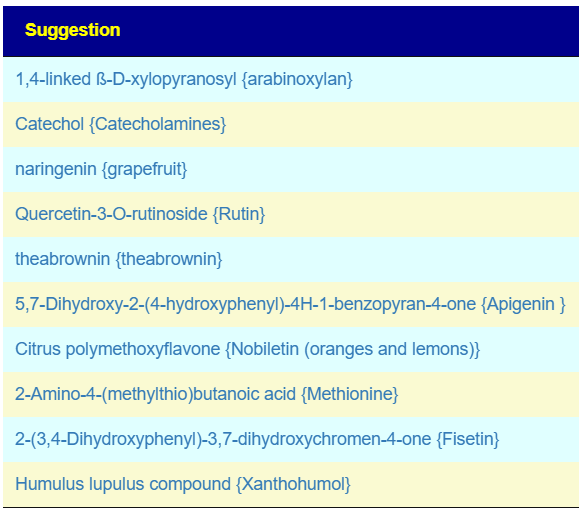

Vitamins

Common and OTC Supplements
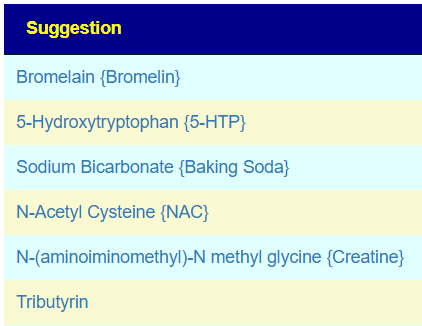
Probiotics x PubMed
This list is done using PubMed studies.

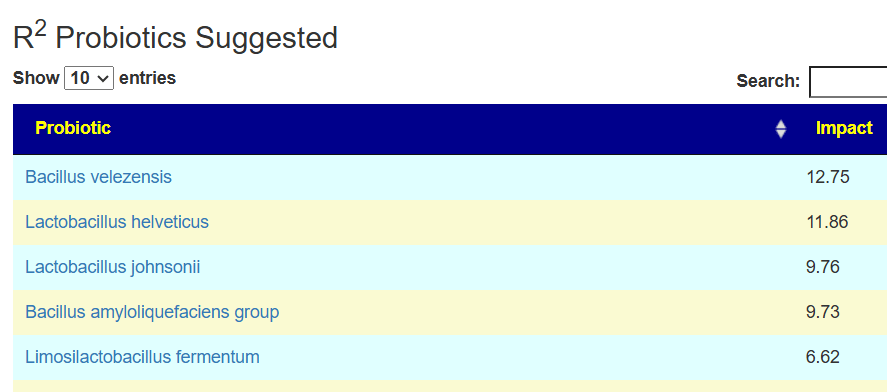
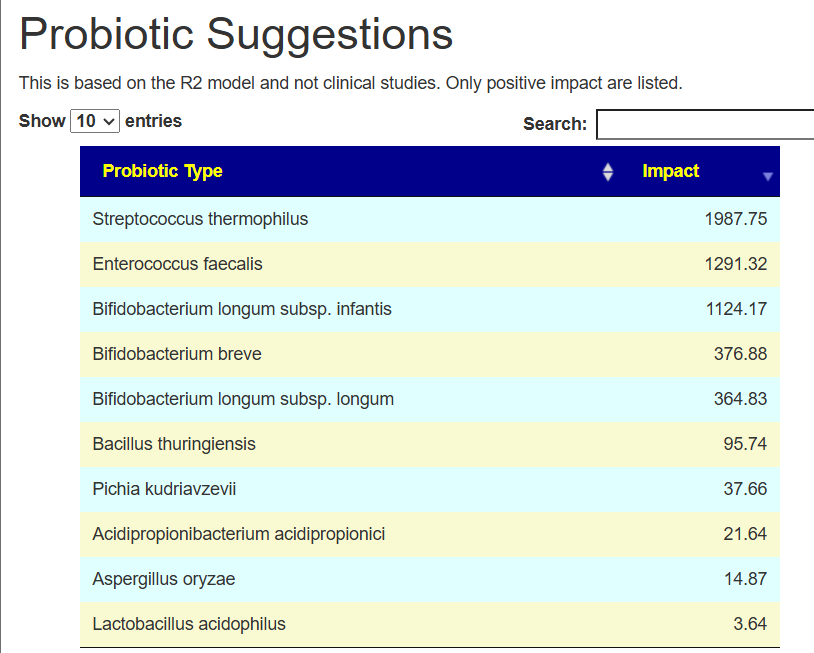
Probiotics x R2 Model
I prefer the R2 Model because we have a lot more data to use than with PubMed. On the flip side, this does not have clinical studies supporting the choices.

The top probiotic Bacillus thuringiensis suitable for human consumption may be a challenge. Most retail products are formulated to control caterpillars, worms, or mosquito larvae in gardens and standing water, not for ingestion or probiotic use.
Non-Human Retail options
- Home Depot sells Monterey B.T. Bacillus Thuringiensis Pint Concentrate as an online retail product.
- Wilco Farm Stores / Farm Store lists Bonide Bt Bacillus Thuringiensis as a garden pest-control product.
- DoMyOwn sells a dedicated Bt category and says it’s available through their store rather than big-box shelves.
- FBN lists Bt ingredient-based products, including Bacillus thuringiensis subspecies tenebrionis.
- DIY Pest Control lists Bt products and notes common trade names like Thuricide and Mosquito Dunks.
Summary
I look forward to see how well this alternative approach performs. It does not focus on the bacteria associated with his 141 symptoms — instead, we focus on shifting to a healthy microbiome profile (with very high statistical significance, p < 0.0001,) I would suggest retesting every 3-4 months to track progress.
Questions And Answers
Q: It’s interesting to see how some Odds Ratio based suggestions match with the Consensus Suggestions, and some vary wildly.
- A: Suggestions are based on bacteria target and available literature. Literature is sparse and often without replication of results
- The safest path would be to start with items that are in agreement.
Q: I had one question with regard to whole milk, dairy, and lactose. The Odds Ratio analysis suggested these were positives – this makes my life a lot easier as I was using milk to help ferment and increase the CFU of the probiotics I used last year and hoped to again, and I eat a fair amount of dairy in general (I mostly eat a vegetarian diet with occasional fish, and dairy helps with my protein intake).
However when I ran the Consensus Suggestions earlier this week I got scores of -294.9 for bovine milk products, -120 for whole cow milk, and -158.6 for lactose.
- A: I favor the Odds Ratio. On this point as you have no issues with dairy, keep to your current usage.
Q: Does this mean I likely need to make a choice between the Consensus Suggestions route (which I followed last year) and the new Odds Ratio route?
- A: No, you could start doing a consensus of the consensus and odds ratio. Then add in items that disagree. I would suggest using an ratio evaluation:
- Consensus: -120 with min of -960, so -(120/960) = -12.5%
- Odds Ratio: 4 with max of 8, so 4/8 =50%
- Thus odds ratio would win
Postscript – and Reminder
Other posts dealing with ME/CFS or Long COVID.
I am not a licensed medical professional, and the laws where I live prohibit any activity that could be interpreted as practicing medicine or giving personal medical advice. My work is limited to academic and analytical models, and I restrict myself to the language of science and statistics rather than clinical recommendations.
I cannot tell anyone what they should or should not take. Instead, I can present information about items that, based on numerical and statistical analysis, appear to have better odds of improving microbiome-related measures. I am a trained, experienced statistician with appropriate degrees and professional affiliations, and my role is to interpret data—not to treat patients.
All information I provide is for educational and informational purposes only and is not a substitute for professional medical advice, diagnosis, or treatment. Any ideas, rankings, or “suggestions” derived from my analyses must be reviewed and approved by your qualified medical professional before you decide to act on them.
The answers and explanations I provide describe my reasoning and methodology. They are not intended as medical advice for you or for anyone else, and they do not create a doctor–patient or provider–patient relationship. Always consult a knowledgeable licensed healthcare professional before starting, changing, or stopping any treatment, supplement, or health-related regimen.