A reader asked me about my hypothesize about methylation and antibiotics. I will respond to the first item in this post. The hypothesis is that DNA impacts methylation and is thus connected to some symptoms.
I have done one post on methylation in the past, Methylation, CFS and the microbiome [2016]. Methylation was a hot research topic reaching a peak in 2015 when interest started to drop according to PubMed
Irritable Bowel Syndrome
- Genome-wide DNA methylation profiling of peripheral blood mononuclear cells in irritable bowel syndrome[2016].
- “Genome-wide DNA methylation profiling of IBS patients compared with HCs identified 133 differentially methylated positions (DMPs) (mean difference ≥10%; p < 0.05).”
- Epigenetic changes in differential methylation of subcommissural organ (SCO)-Spondin (SSPO) gene were positively correlated with hospital anxiety and depression scores in IBS patients (r > 0.4 and false discovery rate <0.05).
- Peptide Therapeutics and the Pharmaceutical Industry: Barriers Encountered Translating from the Laboratory to Patients [2016]. Severe difficulties in using simple solutions — they don’t work!
Fibromyalgia
- ” Epigenome-wide analysis of DNA methylation was conducted…three CPGs reached significant p-values in the replication sample, including malate dehydrogenase 2 (MDH2; p-value 0.017), tetranectin (CLEC3B; p-value 0.039), and heat shock protein beta-6 (HSPB6; p-value 0.016)…we found evidence for the involvement of epigenetic factors. ” [2016]
- “The majority of differentially methylated (DM) sites (91%) were attributable to increased values in the women with FM. The DM sites included significant biological clusters involved in neuron differentiation/nervous system development, skeletal/organ system development, and chromatin compaction. Genes associated with DM sites whose function has particular relevance to FM included BDNF, NAT15, HDAC4, PRKCA, RTN1, and PRKG1.” [2013]
Chronic Fatigue Syndrome
- Epigenetic modifications and glucocorticoid sensitivity in Myalgic Encephalomyelitis/Chronic Fatigue Syndrome (ME/CFS) [2017].
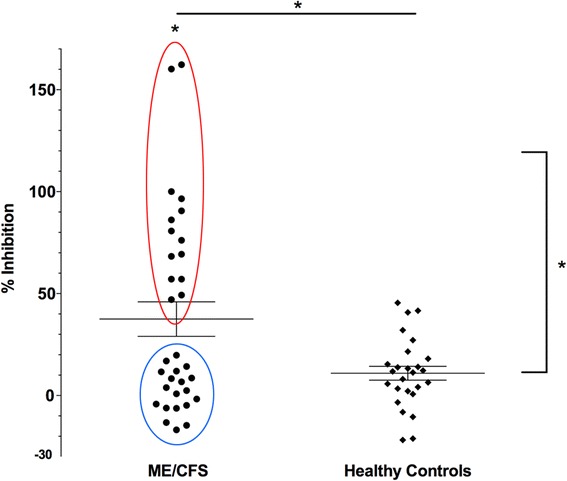
” we report DNA methylation differences in PBMCs of ME/CFS patients, some of which were significantly associated with overall quality of life as well as glucocorticoid hypersensitivity in a subgroup of ME/CFS patients. … we found that genes such as GSTM1, MYO3B, GSTM5, and ATP6V0E2 showed significant epigenetic modifications in ME/CFS. “ - Epigenome-wide DNA methylation patterns associated with fatigue in primary Sjögren’ssyndrome [2016]. “Some genes involved in regulation of the immune system and in inflammation are differently methylated in pSS patients with high vs low fatigue…. the most pronounced hypomethylation in pSS high fatigue annotated to the SBF2-antisense RNA1 gene.”
- “Overall NR3C1-1F DNA methylation was lower in [female] patients with CFS than in controls.” [2015]
- ” We found an increased abundance of differentially methylated genes related to the immune response, cellular metabolism, and kinase activity. Genes associated with immune cell regulation, the largest coordinated enrichment of differentially methylated pathways, showed hypomethylation within promoters and other gene regulatory elements in CFS.” [2014]
Bacteria
While we are dealing with DNA – the changes are epigenetic (relating to or arising from nongenetic influences on gene expression — i.e. the environment, food, stress and most important, the microbiome). This page describes it better.
- The Helicobacter pylori Methylome: Roles in Gene Regulation and Virulence[2017].
- “High-fat diets (HFD) has been reported to induce DNA methylation changes in white adipose tissue” [2017]
- Identification of a Pseudomonas aeruginosa PAO1 DNA Methyltransferase, Its Targets, and Physiological Roles [2017]. A bacteria which can undergo methylization changes (which impacts what gets delivers to the body)
- Pathogenic mechanisms of intracellular bacteria[2017]. “Epigenetic regulators produced by intracellular bacteria alter the epigenotype and gene expression pattern of host cells and play an important role in pathogenesis.”
- Wilson Disease: Epigenetic effects of choline supplementation on phenotype and clinical course in a mouse model [2016]. The impact on increase in one amino acid changed the epigenetic state.
In summary:
“The human microbiota and epigenetic processes have both been shown to play a crucial role in health and disease. However, there is extremely scarce information on epigenetic modulation of microbiota members except for a few pathogens. … It would thus appear likely that such mechanisms are widespread for most bacterial members of the microbiota.” [2016]
Bottom Line
Yes, methylation is a significant factor for a subset of CFS patients, likely more common among female patients than male patients. The problem is epigentic: induced issues (environment). We know very little about what influences these epigentic changes. We know that some amino acids have an impact.
My preferred hypothesis is that the epigenetic shift is caused by a shift of metabolites (chemicals, including amino acids) produced by the microbiome. This would explain why some people with CFS and the same genes do or do not have epigenetic changes influencing methylation. It is preferred also, because it is actionable to correct the root cause! What is the root cause — a microbiome (gut bacteria) shift. Yes, methyl-b12 (1000 mcg/ 1 mg seems to be the effective ongoing dosage, see this post) may help a subset but this is treating the end result and not the root cause.
