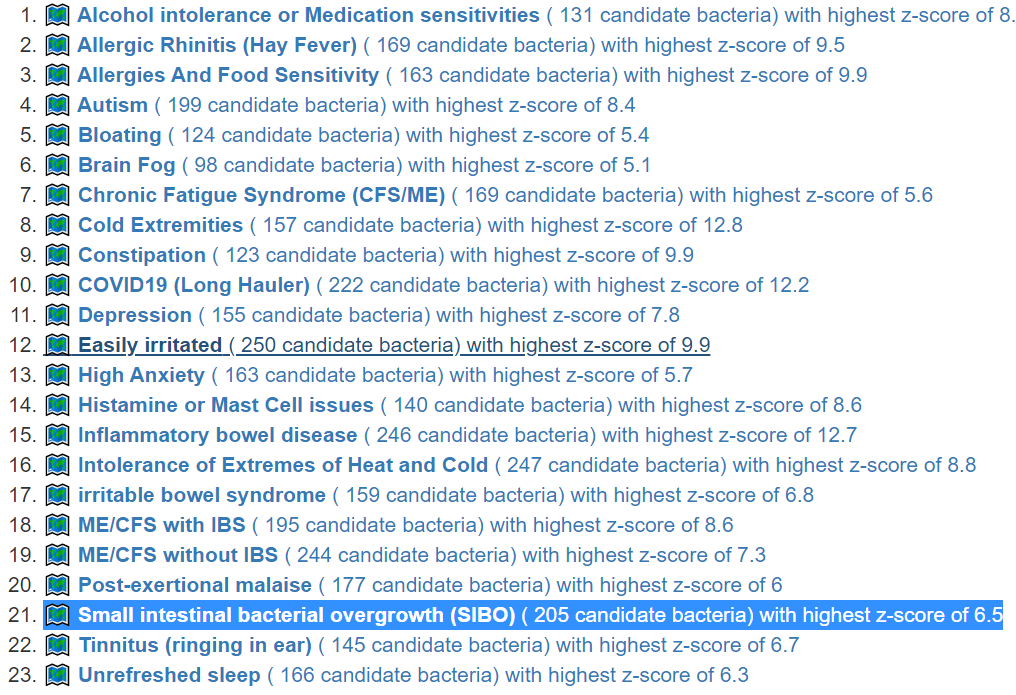
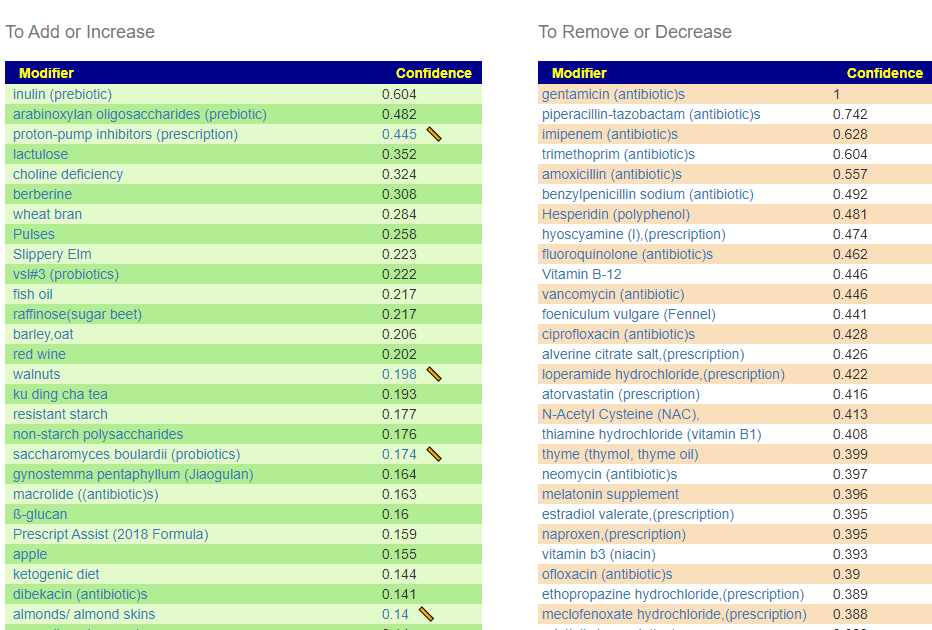
This is a common symptom for many people. This is reported often in samples, and thus being examined if it reaches our threshold for inclusion as defined in A new specialized selection of suggestions links. It does. We are not being specific about the type of constipation. The numbers are less than desired. If you have SIBO and a sample processed thru biomesight (if you have OmbreLab, see how to do this . 📹Video on Transferring Data from Ombre/Thryve to Biomesight ) and then add SIBO as a symptoms. The online data is recomputed once a month so it will improve identification and suggestions.
The second approach is to look at where bacteria identified as statistically significant agrees with KEGG data on Methane, Hydrogen and Hydrogen Sulfide – what is measured in breath tests.
Small intestine aspirate and fluid culture. This is currently the gold standard test for bacterial overgrowth. To obtain the fluid sample, doctors pass a long, flexible tube (endoscope) down your throat and through your upper digestive tract to your small intestine. A sample of intestinal fluid is withdrawn and then tested in a laboratory for the growth of bacteria.
Small intestinal bacterial overgrowth (SIBO) – Mayo Clinic
IMHO, the sample should go thru shotgun processing and not culturing (growth of bacteria)
“Patients with functional dyspepsia also were found to have a greater relative abundance of Streptococcus and decreases in the relative abundance of other genera such as Prevotella, Veillonella, and Actinomyces compared with control subjects, suggesting that their symptoms may be related to alterations of their microbiome at this site…. 16S ribosomal RNA (rRNA) sequencing revealed that SIBO subjects had 4-fold significantly higher relative abundance of Proteobacteria and 1.6-fold significantly lower Firmicutes than non-SIBO subjects. Furthermore, altered Proteobacterial profiles were found that correlated with symptom severity. ”
Current and Future Approaches for Diagnosing Small Intestinal Dysbiosis in Patients With Symptoms of Functional Dyspepsia [2022]
Reservation of SIBO as a Condition
I will be upfront — my motto is Ostende mihi testimonium – Show me the evidence. I do not dispute getting results from Breath Tests – but reading the latest literature [see 2022], it is likely that most of the breath tests are incorrectly done. This is especially true give the type of advice that I have seen on social media compared to the best literature (see above link). SIBO is associated with so many other conditions that I view it as a shared symptom, such as head ache or constipation. Many clinical studies are done in reference to SIBO in the context of this or that condition. This implies that researchers are seeing very different subsets in what is called SIBO. Just with the three existing breath tests, we see 3! i.e. 6 potential subtypes of SIBO – each with different bacteria involved and thus different treatment plan being likely.
In preparing this study, I attempted to disprove my gut feeling but looking for what should be there and could not find it. See below.
Study Populations:
| Symptom | Reference | Study |
| Small intestinal bacterial overgrowth (SIBO) | 1186 | 35 |
- Bacteria Detected with z-score > 2.6: found 205 items, highest value was 6.5
- Enzymes Detected with z-score > 2.6: found 335 items, highest value was 7.9
- Compound Detected with z-score > 2.6: found No items
Interesting Significant Bacteria
All bacteria was found to be low. This may suggests that bacteria that would consume Methane, Hydrogen and Hydrogen Sulfide may be deficient.
| Bacteria | Reference Mean | Study | Z-Score |
| Pectinatus cerevisiiphilus (species) | 197 | 93 | 6.5 |
| Alkaliphilus (genus) | 3463 | 1239 | 6.1 |
| Pectinatus (genus) | 201 | 101 | 6 |
| Alkaliphilus crotonatoxidans (species) | 3424 | 1235 | 5.9 |
| Eubacteriales Family XIII. Incertae Sedis (family) | 441 | 157 | 5.7 |
| Streptococcus fryi (species) | 128 | 52 | 5.6 |
| Thermoanaerobacter (genus) | 60 | 25 | 5.6 |
| Sedimentibacter (genus) | 1397 | 460 | 5.5 |
| Tissierellia incertae sedis (norank) | 1402 | 464 | 5.5 |
| Legionellales (order) | 80 | 41 | 5.2 |
| Legionellaceae (family) | 80 | 41 | 5.2 |
| Legionella (genus) | 80 | 41 | 5.2 |
| Mogibacterium (genus) | 440 | 176 | 5 |
| Adlercreutzia (genus) | 391 | 170 | 5 |
Interesting Enzymes
As with bacteria, all of the enzymes found significant were too low. We have a lot of them!
| Enzyme | Reference Mean | Study Mean | Z-Score |
| propanoate:CoA ligase (AMP-forming) (6.2.1.17) | 1884 | 267 | 7.9 |
| all-trans-zeta-carotene:acceptor oxidoreductase (1.3.99.26) | 2217 | 678 | 6.6 |
| 15-cis-phytoene:acceptor oxidoreductase (neurosporene-forming) (1.3.99.28) | 2217 | 678 | 6.6 |
| 15-cis-phytoene:acceptor oxidoreductase (zeta-carotene-forming) (1.3.99.29) | 2217 | 678 | 6.6 |
| 15-cis-phytoene:acceptor oxidoreductase (lycopene-forming) (1.3.99.31) | 2217 | 678 | 6.6 |
| (2S,3R)-3-hydroxybutane-1,2,3-tricarboxylate pyruvate-lyase (succinate-forming) (4.1.3.30) | 1434 | 295 | 6.6 |
| propanoyl-CoA:oxaloacetate C-propanoyltransferase (thioester-hydrolysing, 1-carboxyethyl-forming) (2.3.3.5) | 1480 | 329 | 6.5 |
| catechol:oxygen 2,3-oxidoreductase (ring-opening) (1.13.11.2) | 3219 | 1059 | 6.4 |
| 5-aminopentanoate:2-oxoglutarate aminotransferase (2.6.1.48) | 2360 | 560 | 6.4 |
| (S)-3-amino-2-methylpropanoate:2-oxoglutarate aminotransferase (2.6.1.22) | 2353 | 555 | 6.4 |
| (S)-2-hydroxyglutarate:quinone oxidoreductase (1.1.5.13) | 2188 | 423 | 6.3 |
| L-carnitinyl-CoA hydro-lyase [(E)-4-(trimethylammonio)but-2-enoyl-CoA-forming] (4.2.1.149) | 1445 | 225 | 6.2 |
| glutarate, 2-oxoglutarate:oxygen oxidoreductase ((S)-2-hydroxyglutarate-forming) (1.14.11.64) | 1298 | 221 | 6.2 |
| biuret amidohydrolase (3.5.1.84) | 778 | 266 | 6.1 |
| L-carnitine,NAD(P)H:oxygen oxidoreductase (trimethylamine-forming) (1.14.13.239) | 1189 | 218 | 6.1 |
| medium-chain acyl-CoA:electron-transfer flavoprotein 2,3-oxidoreductase (1.3.8.7) | 1285 | 226 | 6.1 |
| succinyl-CoA:3-oxo-acid CoA-transferase (2.8.3.5) | 1289 | 246 | 6 |
| n/a (3.4.14.13) | 1253 | 215 | 6 |
| (2S,3S)-2-hydroxybutane-1,2,3-tricarboxylate hydro-lyase [(Z)-but-2-ene-1,2,3-tricarboxylate-forming] (4.2.1.79) | 1297 | 312 | 6 |
| RNA-3′-phosphate:RNA ligase (cyclizing, AMP-forming) (6.5.1.4) | 1218 | 225 | 5.9 |
| D-glucose:NAD(P)+ 1-oxidoreductase (1.1.1.47) | 1237 | 222 | 5.9 |
| n/a (3.4.23.49) | 1306 | 247 | 5.9 |
| (2R)-2-O-phospho-3-sulfolactate hydrogen-sulfite-lyase (phosphoenolpyruvate-forming) (4.4.1.19) | 1488 | 217 | 5.9 |
| CDP-ribitol:4-O-di[(2R)-1-glycerophospho]-N-acetyl-beta-D-mannosaminyl-(1->4)-N-acetyl-alpha-D-glucosaminyl-diphospho-ditrans,octacis-undecaprenol ribitolphosphotransferase (2.7.8.14) | 2268 | 671 | 5.9 |
| CDP-ribitol:4-O-[1-D-ribitylphospho-(2R)-1-glycerophospho]-N-acetyl-beta-D-mannosaminyl-(1->4)-N-acetyl-alpha-D-glucosaminyl-diphospho-ditrans,octacis-undecaprenol ribitolphosphotransferase (2.7.8.47) | 2268 | 671 | 5.9 |
| UDP-N-acetyl-alpha-D-glucosamine:lipopolysaccharide N-acetyl-D-glucosaminyltransferase (2.4.1.56) | 1054 | 181 | 5.8 |
| D-galactaro-1,4-lactone lyase (ring-opening) (5.5.1.27) | 1676 | 380 | 5.7 |
| hydrogen-sulfide:flavocytochrome c oxidoreductase (1.8.2.3) | 1145 | 88 | 5.6 |
| 3-methylcrotonoyl-CoA:carbon-dioxide ligase (ADP-forming) (6.4.1.4) | 1099 | 135 | 5.6 |
| [SoxY protein]-S-sulfosulfanyl-L-cysteine sulfohydrolase (3.1.6.20) | 1164 | 86 | 5.6 |
| (R)-lactate hydro-lyase (4.2.1.130) | 1550 | 290 | 5.6 |
| CTP:5,7-diacetamido-3,5,7,9-tetradeoxy-L-glycero-alpha-L-manno-nonulosonic acid cytidylyltransferase (2.7.7.81) | 1121 | 117 | 5.4 |
| sulfite:oxygen oxidoreductase (1.8.3.1) | 1180 | 82 | 5.4 |
| L-kynurenine hydrolase (3.7.1.3) | 989 | 98 | 5.4 |
| glutaryl-CoA:electron-transfer flavoprotein 2,3-oxidoreductase (decarboxylating) (1.3.8.6) | 1230 | 212 | 5.4 |
| L-carnitine:CoA ligase (AMP-forming) (6.2.1.48) | 1376 | 442 | 5.4 |
| (1E,3E)-4-hydroxybuta-1,3-diene-1,2,4-tricarboxylate 1,2-hydro-lyase (2-hydroxy-4-oxobutane-1,2,4-tricarboxylate-forming) (4.2.1.83) | 1038 | 102 | 5.3 |
| alkane,reduced-rubredoxin:oxygen 1-oxidoreductase (1.14.15.3) | 1068 | 103 | 5.2 |
| glutaredoxin:hydroperoxide oxidoreductase (1.11.1.25) | 988 | 107 | 5.2 |
| n/a (3.4.21.96) | 7235 | 4398 | 5.2 |
| isoquinoline:acceptor 1-oxidoreductase (hydroxylating) (1.3.99.16) | 987 | 112 | 5.2 |
| sn-glycerol-3-phosphate:oxygen 2-oxidoreductase (1.1.3.21) | 472 | 112 | 5.2 |
| reduced coenzyme F420:NADP+ oxidoreductase (1.5.1.40) | 1751 | 594 | 5.2 |
| 4-fumarylacetoacetate fumarylhydrolase (3.7.1.2) | 984 | 124 | 5.1 |
| [sulfatase]-L-cysteine:oxygen oxidoreductase (3-oxo-L-alanine-forming) (1.8.3.7) | 1014 | 215 | 5.1 |
| S-methyl-5′-thioadenosine:phosphate S-methyl-5-thio-alpha-D-ribosyl-transferase (2.4.2.28) | 3707 | 1421 | 5.1 |
| gamma-butyrobetainyl-CoA:electron-transfer flavoprotein 2,3-oxidoreductase (1.3.8.13) | 1398 | 456 | 5.1 |
| (2->6)-beta-D-fructan fructanohydrolase (3.2.1.65) | 189 | 43 | 5.1 |
| (S)-lactate:oxygen 2-oxidoreductase (1.1.3.2) | 662 | 219 | 5.1 |
| (E)-4-(trimethylammonio)but-2-enoyl-CoA:L-carnitine CoA-transferase (2.8.3.21) | 1384 | 458 | 5.1 |
| benzoyl-CoA,NADPH:oxygen oxidoreductase (2,3-epoxydizing) (1.14.13.208) | 107 | 32 | 5.1 |
| (S)-mandelate:acceptor 2-oxidoreductase (1.1.99.31) | 112 | 34 | 5.1 |
| 5-dehydro-4-deoxy-D-glucarate hydro-lyase (decarboxylating; 2,5-dioxopentanoate-forming) (4.2.1.41) | 972 | 100 | 5.1 |
| 2,5-dioxopentanoate:NADP+ 5-oxidoreductase (1.2.1.26) | 978 | 127 | 5 |
| ferulate:CoA ligase (ATP-hydrolysing) (6.2.1.34) | 973 | 97 | 5 |
| homogentisate:oxygen 1,2-oxidoreductase (ring-opening) (1.13.11.5) | 985 | 116 | 5 |
| ADP-glucose:D-glycerate 2-alpha-D-glucosyltransferase (2.4.1.268) | 67 | 24 | 5 |
| N,N-dimethylaniline,NADPH:oxygen oxidoreductase (N-oxide-forming) (1.14.13.8) | 1030 | 247 | 5 |
| ATP:[protein]-N6-D-ribulosyl-L-lysine 3-phosphotransferase (2.7.1.172) | 1095 | 80 | 5 |
Significant Bacteria and KEGG modelling
To me, this is where the exploration could get interesting.
The following are consumers of these three compound according to KEGG and are statistically significant in our study, Those marked with * also produces some of these compounds. Whether a bacteria is producing or consuming may depend on what is in the microbiome environment (i.e. enzyme quantity).
- Adlercreutzia equolifaciens *
- Butyrivibrio proteoclasticus *
- Corynebacterium aurimucosum *
- Emticicia oligotrophica *
- Escherichia albertii *
- Escherichia coli *
- Haemophilus parahaemolyticus *
- Phascolarctobacterium faecium *
- Phocaeicola salanitronis *
- Pseudobutyrivibrio xylanivorans
- Ruminococcus albus *
- Streptococcus anginosus
- Streptococcus equinus
- Streptococcus oralis (A.K.A. Streptococcus dentisani)
- Streptococcus vestibularis
- Turicibacter sanguinis
My perspective is that the consumption (and production) of Methane, Hydrogen and Hydrogen Sulfide is dependent on the availability of enzymes and chemicals produced by other bacteria. Those are not measured in testing — only those three easy to test by last millennium lab tests.
Of the items above, two are potentially available as retail probiotics now, or soon:
- Streptococcus dentisani, studies: [2020] [2021] [2019] [2018]
- Escherichia coli (Mutaflor, Symbioflor-2 (US Source, World Wide Source) )
Concerning the enzymes listed above, I will leave it to the reader to see where they occur in the following example of a KEGG metabolism diagram.
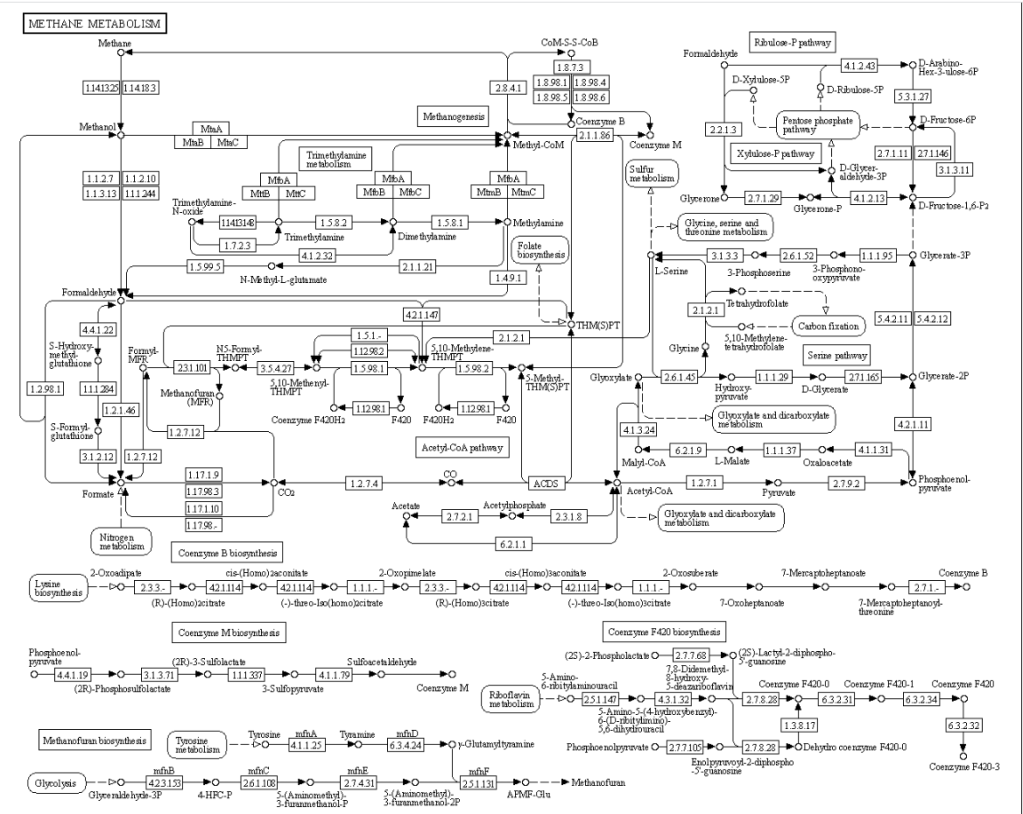
I checked PubMed if there were any studies that used KEGG data for SIBO, there was just one: Association of Differential Metabolites With Small Intestinal Microflora and Maternal Outcomes in Subclinical Hypothyroidism During Pregnancy [2022] – “KEGG pathway analysis revealed that differential metabolites were mainly involved in bile secretion, cholesterol metabolism, and other pathways”
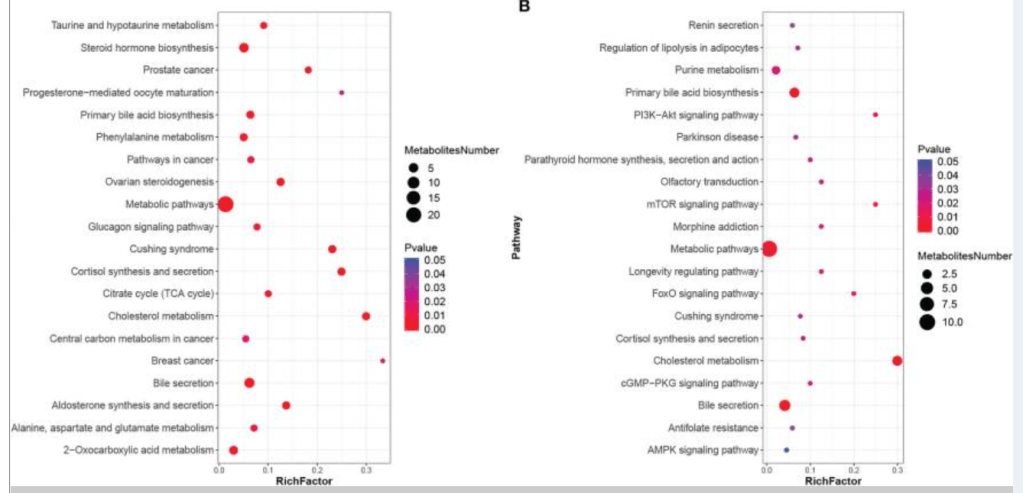
What I find really irritating is that some of the bacteria inferred to cause SIBO lacks the genetic/genes/enzymes to produce Methane, Hydrogen or Hydrogen Sulfide. My sole comment is last millennium medical belief versus current millennium medical fact.
Bottom Line
SIBO is difficult to treat. I suspect one of the cause is going with a simplistic naïve view of this condition: It is caused by having too many producers of Methane, Hydrogen and Hydrogen Sulfide. In this study, we found the opposite — it is an absence of consumers of Methane, Hydrogen and Hydrogen Sulfide. Furthermore, the existing consumers may be inhibited in this function by the absence of enzymes needed to consume. This data is coming from downstream, so it may not apply apply fully to the small intestine – however the chemicals and enzymes flowing from the small intestine would impact a stool sample.
If the naïve view was correct, the specific bacteria would have been identified decades ago and SIBO would be resolved by a round of antibiotics targeted at them. That is not people’s experience dealing with SIBO.
The bottom citation is long and well worth reading. It covers issues well with SIBO testing.
Coincident with advances in medical science, diagnostic testing evolved from small bowel culture to breath tests and on to next-generation, culture-independent microbial analytics. The advent and ready availability of breath tests generated a dramatic expansion in both the rate of diagnosis of SIBO and the range of associated gastrointestinal and nongastrointestinal clinical scenarios. However, issues with the specificity of these same breath tests have clouded their interpretation and aroused some skepticism regarding the role of SIBO in this expanded clinical repertoire.
Small Intestinal Bacterial Overgrowth-Pathophysiology and Its Implications for Definition and Management [2022]
Measurement of breath hydrogen (H2) and methane (CH4) excretion after ingestion of test‐carbohydrates is used for different diagnostic purposes. There is a lack of standardization among centers performing these tests and this, together with recent technical developments and evidence from clinical studies, highlight the need for a European guideline.
European guideline on indications, performance, and clinical impact of hydrogen and methane breath tests in adult and pediatric patients: European Association for Gastroenterology, Endoscopy and Nutrition, European Society of Neurogastroenterology and Motility, and European Society for Paediatric Gastroenterology Hepatology and Nutrition consensus [2022]
… SIBO has been associated with multiple conditions including IBS, rosacea, hepatic encephalopathy, obesity, gastroparesis, Parkinson’s disease, fibromyalgia, chronic pancreatitis, end‐stage renal disease, and inflammatory bowel diseases.
H2BT has become the most used test for SIBO in clinical practice; however, this is largely due to its ease of use, non‐invasive character and low cost and not based on evidence from clinical trials.
In short, I do not know what causes the breath test results — there is an abundance of speculation, often held to as fact. There is a absence of solid evidence. What I do know is that bacteria moves up and down in the body – think of the analogy of salmon which moves upstream against strong currents. We see strong statistical shifts in stool samples as a whole, this is evidence that hints that correction may have as a side effect, relief of SIBO.