Some smart readers wrote saying that they are familiar with R but not the microbiome. This post is intended to kickstart them on a simple desired goal:
Which bacteria (or combination of bacteria) are associated with certain symptoms.
We have 240 symptoms and thousands of bacteria … oops, a big problem. We do not have a lot of microbiomes with symptoms list — so we are likely forced to deal with only the most common symptoms at present.
Information about bacteria is layered
A quick read on wikipedia on bacteria taxonomy may help. If you just grab the Taxonomy file, you will see the species counted in with it’s parent genus, who is counted in with family, etc… not the best scenario for analysis.

To simplify out matters, I have done extracts on each layer and dropped them at http://lassesen.com/ubiome/

So to load one layer, it’s just a few lines
library(tidyverse)
Species <-read_csv(“http://lassesen.com/ubiome/TaxonomyByspecies.csv”)
symptoms <-read_csv(“http://lassesen.com/ubiome/Symptoms.csv” )
samplesymptom <-read_csv(“http://lassesen.com/ubiome/SampleSymptom.csv”)
samplesWithTitle <- inner_join(samplesymptom, symptoms, by=”symptomId” )
Looking at this one, we find some 2,181 columns (species) and a sampleId (that matches that in symptoms).

From my prior post, you can see how we can get the most common symptom (symptomId=1,”General: Fatigue”) and then a list of sampleId with:
HasFatigue <-samplesWithTitle %>% filter(symptomId==1)

We need to only include observations that reported ANY symptoms, which is easy to do:
PeopleWithSymptoms <- samplesymptom %>% group_by(sampleId) %>% summarize(peopleWith=n()) %>% arrange(peopleWith)
Next we joins this with the level of taxonomy we wish to explore.
SpeciesWithSymptoms <-merge(PeopleWithSymptoms ,Species, by=”sampleId”)

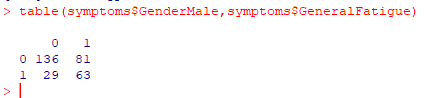
We next put the sets together and set symptomId to 0 for those without fatigue. We have 47.5% of the records with Fatigue.
fulldata <- left_join(SpeciesWithSymptoms,HasFatigue,by=”sampleId”)
fulldata$symptomId =ifelse(is.na(fulldata$symptomId),0,fulldata$symptomId)
#We need to remove the symptom_name
drops <- c(“symptom_name”)
regdata <-fulldata[, !(names(fulldata) %in% drops)]
Picking the Model
I went with a simple linear model for the sake of illustration and toss everything at it!
test <- lm(symptomId ~ .,regdata)
We end up getting a rather long report back:

Where there is a + sign, it indicates increased risk of fatigue as the numbers of bacteria increases, -sign decreased risk of fatigue as the numbers of bacteria increases.
I can now redo it, picking just a few bacteria:

and then looking at the statistics, we see that there is no statistical significance for this relationship (we want to see a p-value < 0.001)

That’s it. No revelations — just an illustration of walking the path. All of the code for above is below.
library(tidyverse)
Species <-read_csv(“http://lassesen.com/ubiome/TaxonomyByspecies.csv”) symptoms <-read_csv(“http://lassesen.com/ubiome/Symptoms.csv” ) samplesymptom <-read_csv(“http://lassesen.com/ubiome/SampleSymptom.csv”) samplesWithTitle <- inner_join(samplesymptom, symptoms, by=”symptomId” ) HasFatigue <-samplesWithTitle %>% filter(symptomId==1)
PeopleWithSymptoms <- samplesymptom %>% group_by(sampleId) %>% summarize(peopleWith=n()) %>% arrange(peopleWith)
SpeciesWithSymptoms <-merge(PeopleWithSymptoms ,Species, by=”sampleId”)
fulldata <- left_join(SpeciesWithSymptoms,HasFatigue,by=”sampleId”)
fulldata$symptomId =ifelse(is.na(fulldata$symptomId),0,fulldata$symptomId)
drops <- c(“symptom_name”)
regdata <-fulldata[, !(names(fulldata) %in% drops)]
test <- lm(symptomId ~ .,regdata)
Doing a run with Phylum,
Phylum <-read_csv(“http://lassesen.com/ubiome/TaxonomyByPhylum.csv”) symptoms <-read_csv(“http://lassesen.com/ubiome/Symptoms.csv” ) samplesymptom <-read_csv(“http://lassesen.com/ubiome/SampleSymptom.csv”) samplesWithTitle <- inner_join(samplesymptom, symptoms, by=”symptomId” ) HasFatigue <-samplesWithTitle %>% filter(symptomId==1)
PeopleWithSymptoms <- samplesymptom %>% group_by(sampleId) %>% summarize(peopleWith=n()) %>% arrange(peopleWith)
PhylumWithSymptoms <-merge(PeopleWithSymptoms ,Phylum, by=”sampleId”)
fulldata <- left_join(PhylumWithSymptoms,HasFatigue,by=”sampleId”)
fulldata$symptomId =ifelse(is.na(fulldata$symptomId),0,fulldata$symptomId)
drops <- c(“symptom_name”)
regdata <-fulldata[, !(names(fulldata) %in% drops)]
test <- lm(symptomId ~ .,regdata)
We find statistical significance with 45% explained by the phylum.