FYI: Bug in http://microbiomeprescription.azurewebsites.net/ OtherLabs Analysis.
I appear to have flip a sign during my rework. last weekend. So Take and Avoid are flipped.
It will be corrected this weekend.
FYI: Bug in http://microbiomeprescription.azurewebsites.net/ OtherLabs Analysis.
I appear to have flip a sign during my rework. last weekend. So Take and Avoid are flipped.
It will be corrected this weekend.
A reader asked for a comparison between various labs (apart from ubiome).
The table is below (also available here as a PDF file)
CompareLabs
| Row Labels | Bioscreen | Biovis Microbiome Plus | Diagnostic Solution GI-Map | Genova Gi Effects | Genova Parasitology | Kyber Kompakt | Medivere: Darm Mikrobiom Stuhltest | Medivere: Darn Magen Diagnostik | Medivere: Gesundsheitscheck Darm | Metagenomics Stool (De Meirleir) | Smart Gut | Verisana | Viome | Grand Total |
| Acetanaerobacterium | 1 | 1 | ||||||||||||
| Acetivibrio | 1 | 1 | ||||||||||||
| Acidaminococcus | 1 | 1 | ||||||||||||
| Acinetobacter | 1 | 1 | ||||||||||||
| Actinobacteria | 1 | 1 | 1 | 3 | ||||||||||
| Actinomyces | 1 | 1 | ||||||||||||
| Adlercreutzia | 1 | 1 | 2 | |||||||||||
| Akkermansia muciniphila | 1 | 1 | 1 | 1 | 4 | |||||||||
| Alistipes | 1 | 1 | 1 | 1 | 4 | |||||||||
| Anaerofustis | 1 | 1 | ||||||||||||
| Anaerostipes | 1 | 1 | ||||||||||||
| Anaerotruncus colihominis | 1 | 1 | 2 | |||||||||||
| Asaccharobacter | 1 | 1 | ||||||||||||
| Atopobium | 1 | 1 | ||||||||||||
| Bacillus | 1 | 1 | ||||||||||||
| Bacteroides | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 10 | |||
| Bacteroides fragilis | 1 | 1 | 1 | 3 | ||||||||||
| Bacteroides intestinalis | 1 | 1 | ||||||||||||
| Bacteroides ovatus | 1 | 1 | ||||||||||||
| Bacteroides vulgatus | 1 | 1 | 2 | |||||||||||
| Bacteroidetes | 1 | 1 | 2 | |||||||||||
| Barnesiella | 1 | 1 | 1 | 3 | ||||||||||
| Bifidobacterium | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 12 | |
| Bifidobacterium longum | 1 | 1 | ||||||||||||
| Bilophila wadsworthia | 1 | 1 | ||||||||||||
| Blautia | 1 | 1 | ||||||||||||
| Butyricicoccus | 1 | 1 | ||||||||||||
| Butyrivibrio | 1 | 1 | ||||||||||||
| Butyrivibrio crossotus | 1 | 1 | 1 | 3 | ||||||||||
| Campylobacter | 1 | 1 | 2 | |||||||||||
| Catenibacterium | 1 | 1 | ||||||||||||
| Christensenellaceae | 1 | 1 | ||||||||||||
| Citrobacter | 1 | 1 | 1 | 1 | 4 | |||||||||
| Citrobacter freundii | 1 | 1 | 2 | |||||||||||
| Clostridia | 1 | 1 | 2 | |||||||||||
| Clostridium | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 7 | ||||||
| Clostridium butyricum | 1 | 1 | ||||||||||||
| Clostridium perfringens | 1 | 1 | ||||||||||||
| Clostridium sporogenes | 1 | 1 | ||||||||||||
| Collinsella | 1 | 1 | 2 | |||||||||||
| Collinsella aerofaciens | 1 | 1 | ||||||||||||
| Coprobacillus | 1 | 1 | ||||||||||||
| Coprococcus | 1 | 1 | 2 | |||||||||||
| Coprococcus eutactus | 1 | 1 | ||||||||||||
| Cyanobacteria | 1 | 1 | ||||||||||||
| Desulfovibrio piger | 1 | 1 | 1 | 3 | ||||||||||
| Dialister | 1 | 1 | ||||||||||||
| Dialister invisus | 1 | 1 | 2 | |||||||||||
| Dorea | 1 | 1 | 2 | |||||||||||
| Eggerthella | 1 | 1 | ||||||||||||
| Eggerthella lenta | 1 | 1 | ||||||||||||
| Enterobacter | 1 | 1 | 1 | 3 | ||||||||||
| Enterobacteriaceae | 1 | 1 | ||||||||||||
| Enterococcaceae | 1 | 1 | ||||||||||||
| Enterococcus | 1 | 1 | 1 | 1 | 1 | 1 | 6 | |||||||
| Erysipelatoclostridium ramosum | 1 | 1 | ||||||||||||
| Escherichia | 1 | 1 | 1 | 3 | ||||||||||
| Escherichia coli | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 8 | |||||
| Escherichia coli Biovare | 1 | 1 | 2 | |||||||||||
| Ethanoligenens | 1 | 1 | 2 | |||||||||||
| Eubacterium | 1 | 1 | 1 | 1 | 1 | 5 | ||||||||
| Eubacterium hallii | 1 | 1 | ||||||||||||
| Euryarchaeota | 1 | 1 | ||||||||||||
| Faecalibacterium | 1 | 1 | ||||||||||||
| Faecalibacterium prausnitzii | 1 | 1 | 1 | 1 | 4 | |||||||||
| Faecalitalea | 1 | 1 | ||||||||||||
| Firmicutes | 1 | 1 | 1 | 3 | ||||||||||
| Fusobacteria | 1 | 1 | 2 | |||||||||||
| Fusobacterium | 1 | 1 | 1 | 3 | ||||||||||
| Gordonibacter | 1 | 1 | ||||||||||||
| Haemophilus | 1 | 1 | ||||||||||||
| Hafnia | 1 | 1 | 2 | |||||||||||
| Hafnia alvei | 1 | 1 | ||||||||||||
| Holdemania | 1 | 1 | ||||||||||||
| Howardella | 1 | 1 | ||||||||||||
| Klebsiella | 1 | 1 | 1 | 1 | 1 | 1 | 6 | |||||||
| Klebsiella pneumoniae | 1 | 1 | 1 | 3 | ||||||||||
| Kluyvera | 1 | 1 | 2 | |||||||||||
| Lachnoclostridium | 1 | 1 | ||||||||||||
| Lactobacillus | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 12 | |
| Lactococcus | 1 | 1 | ||||||||||||
| Lactonifactor | 1 | 1 | ||||||||||||
| Leuconostoc | 1 | 1 | ||||||||||||
| Megamonas | 1 | 1 | ||||||||||||
| Megasphaera | 1 | 1 | ||||||||||||
| Methanobrevibacter smithii | 1 | 1 | 2 | |||||||||||
| Morganella | 1 | 1 | 1 | 3 | ||||||||||
| Morganella morganii | 1 | 1 | ||||||||||||
| Moryella | 1 | 1 | ||||||||||||
| Odoribacter | 1 | 1 | 1 | 1 | 4 | |||||||||
| Olsenella | 1 | 1 | ||||||||||||
| Oscillibacter | 1 | 1 | 2 | |||||||||||
| Oxalobacter formigenes | 1 | 1 | 1 | 3 | ||||||||||
| Papillibacter | 1 | 1 | ||||||||||||
| Parabacteroides | 1 | 1 | 1 | 3 | ||||||||||
| Peptoclostridium | 1 | 1 | ||||||||||||
| Peptoclostridium difficile | 1 | 1 | ||||||||||||
| Prevotella | 1 | 1 | 1 | 1 | 4 | |||||||||
| Proteobacteria | 1 | 1 | 1 | 3 | ||||||||||
| Proteus | 1 | 1 | 1 | 1 | 1 | 1 | 6 | |||||||
| Proteus mirabilis | 1 | 1 | ||||||||||||
| Proteus vulgaris | 1 | 1 | ||||||||||||
| Providencia | 1 | 1 | 2 | |||||||||||
| Pseudoflavonifractor | 1 | 1 | ||||||||||||
| Pseudomonas | 1 | 1 | 1 | 1 | 1 | 5 | ||||||||
| Pseudomonas aeruginosa | 1 | 1 | ||||||||||||
| Roseburia | 1 | 1 | 1 | 1 | 1 | 1 | 6 | |||||||
| Rothia | 1 | 1 | ||||||||||||
| Ruminococcus | 1 | 1 | 1 | 1 | 1 | 1 | 6 | |||||||
| Ruminococcus albus | 1 | 1 | ||||||||||||
| Serratia | 1 | 1 | 1 | 3 | ||||||||||
| Slackia | 1 | 1 | 2 | |||||||||||
| Sporobacter | 1 | 1 | ||||||||||||
| Staphylococcus | 1 | 1 | 2 | |||||||||||
| Streptococcus | 1 | 1 | 1 | 1 | 1 | 5 | ||||||||
| Streptococcus anginosus group | 1 | 1 | ||||||||||||
| Streptococcus parasanguinis | 1 | 1 | ||||||||||||
| Streptococcus salivarius | 1 | 1 | ||||||||||||
| Subdoligranulum | 1 | 1 | ||||||||||||
| Sulfuricurvum | 1 | 1 | ||||||||||||
| Sutterella | 1 | 1 | ||||||||||||
| Syntrophococcus | 1 | 1 | ||||||||||||
| Syntrophus | 1 | 1 | ||||||||||||
| Tannerella | 1 | 1 | ||||||||||||
| Tenericutes | 1 | 1 | ||||||||||||
| Tistrella | 1 | 1 | ||||||||||||
| Turicibacter | 1 | 1 | ||||||||||||
| Veillonella | 1 | 1 | 1 | 3 | ||||||||||
| Verrucomicrobia | 1 | 1 | 2 | |||||||||||
| Xylanibacterium | 1 | 1 | ||||||||||||
| Grand Total | 17 | 42 | 14 | 27 | 7 | 11 | 16 | 16 | 17 | 52 | 23 | 11 | 30 |
A reader forwarded a copy of their Viome report and I have created an entry form for it. It is at: http://microbiomeprescription.com/Kyber/Viome
The report looks like this:

I assume that “Activity” is what the person has.
The report also recommends some food (without providing studies backing them 😦 )
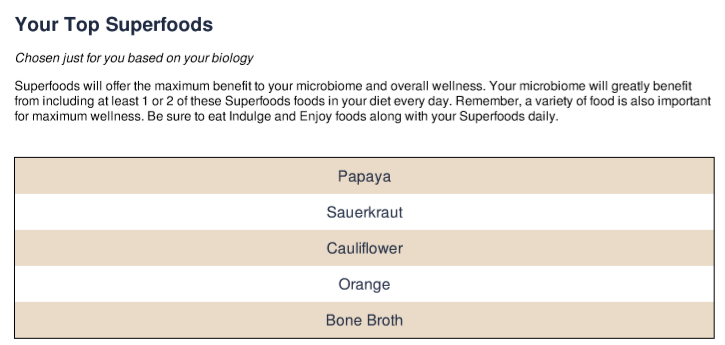
The top recommendation from our site is below. Note that only certain lactobacillus are recommended.

which goes on for several pages. Several items were listed which my analysis suggest should be avoided (barley, almonds, etc) My general impression is their customized for the individual food list is a general healthy living list with perhaps a few minor changes.
I will be researching papaya, cauliflower, orange and bone broth on PubMed to see if there are studies. If they are, I will add them to our database.
Both the other Labs Analysis and Ubiome recommendations have had the revised algorithm applied.
A higher number means more and more studies have found that it has modified the bacteria.
The default recommendations look at:
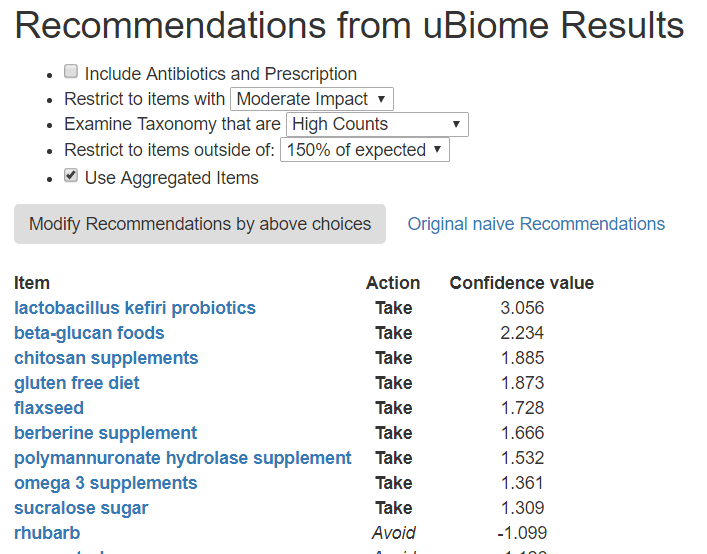
If there are too many items, you may wish to change it to High Impact
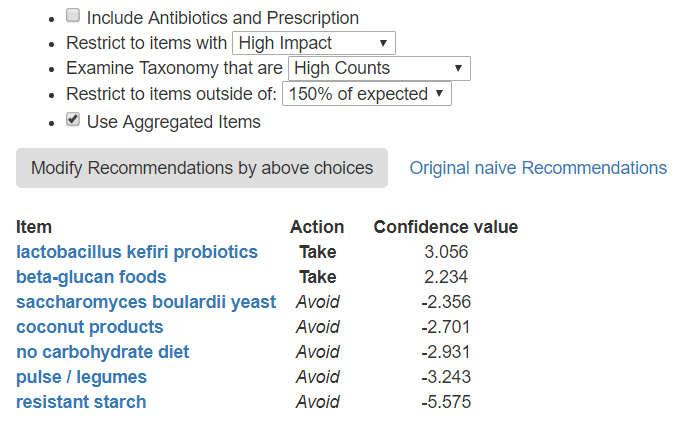
If you are concerned about UNDER GROWTH..
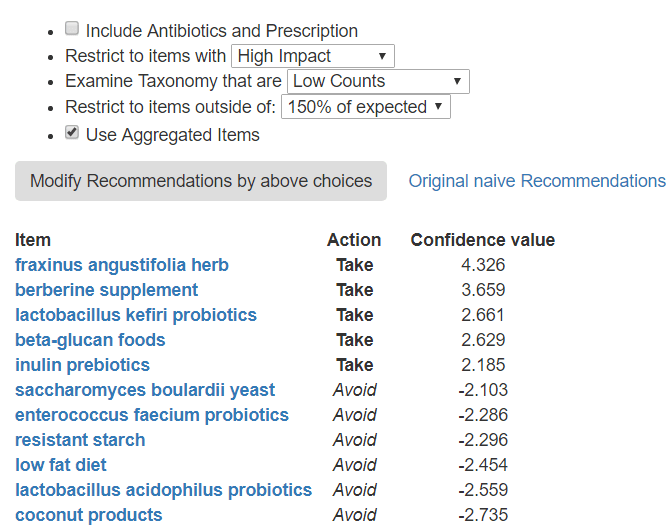
Note that there can be some agreements between the High and Low. For example: avoid coconut products and saccharomyces are on both lists.
If you are interested in an item like ‘beta-glucan foods’, just click on it to see the break down.

This revision should produce better recommendations and be easier to understand. It also provides a tool for those who are concerned about under growths. It also allows people to explore different approaches.
This is an education post to facilitate discussing this approach with your medical professionals. It is not medical advice for the treatment of any medical condition. Always consult with your medical professional before doing any changes of diet, supplements or activity. Some items cites may interfere with prescription medicines.
Another portion of the revision has been completed. This consists of two parts:
I have expanded the reference to every level of the hierarchy

An example is shown below
Click the appropriate link if you have an overgrowth or undergrowth
| Bacteria (% of samples with it) | Modifiers | Data Punk | NCBI Taxonomy |
|---|---|---|---|
| Acinetobacter calcoaceticus/baumannii complex (0.6% of Samples) | Modifiers | Bacteria Information | Literature |
| Bacillus subtilis group (3.59% of Samples) | Modifiers | Bacteria Information | Literature |
| Enterobacter cloacae complex (1.8% of Samples) | Modifiers | Bacteria Information | Literature |
| Lactobacillus casei group (7.78% of Samples) | Modifiers | Bacteria Information | Literature |
| Pseudomonas aeruginosa group (3.59% of Samples) | Modifiers | Bacteria Information | Literature |
| Pseudomonas fluorescens (0% of Samples) | Modifiers | Bacteria Information | Literature |
| Streptococcus anginosus group (1.8% of Samples) | Modifiers | Bacteria Information | Literature |
| Streptococcus dysgalactiae group (1.2% of Samples) | Modifiers | Bacteria Information | Literature |
The literature link takes you to the National Institute of Health site, where you have many additional links of a technical nature.
And DataPunk for the other
If you go to Bacteria Hierarchy or Bacteria Hierarchy Down To Species you will see the percentage of gut samples that have a specific bacteria.
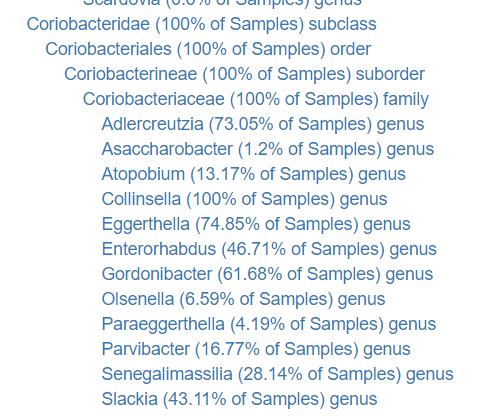

This is important because my long standing question is which low or no count bacteria should we be concerned about to increase. A low Olsenella level is likely not a concern, because most people have none of it. A low Coriobacteriaceae level may be a concern.
If you have a very high level of Olsenella, then this may be a concern. The pending refactor of recommendations will allow have a default (my own assumptions) filtering and the ability for readers to modify it to suit their own assumptions.
These enhancements allow easy access to additional technical information and also allow us to see which bacteria we expect to see normally. This means we can include bacteria that should be there but which are at an abnormally low level in the revised recommendations.