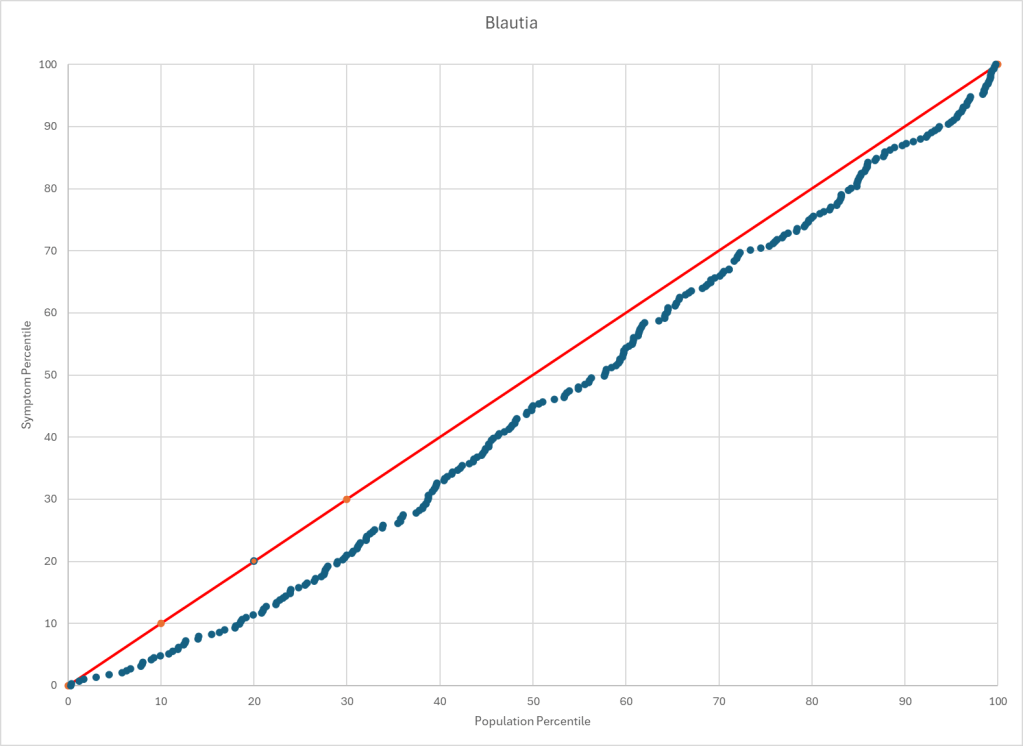
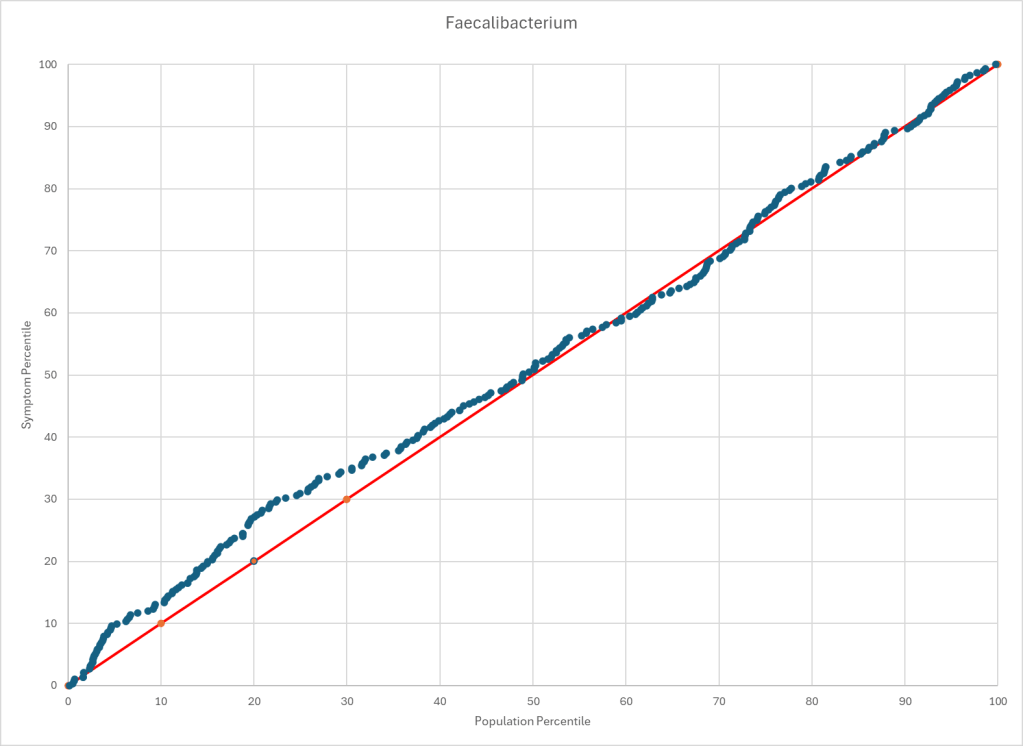
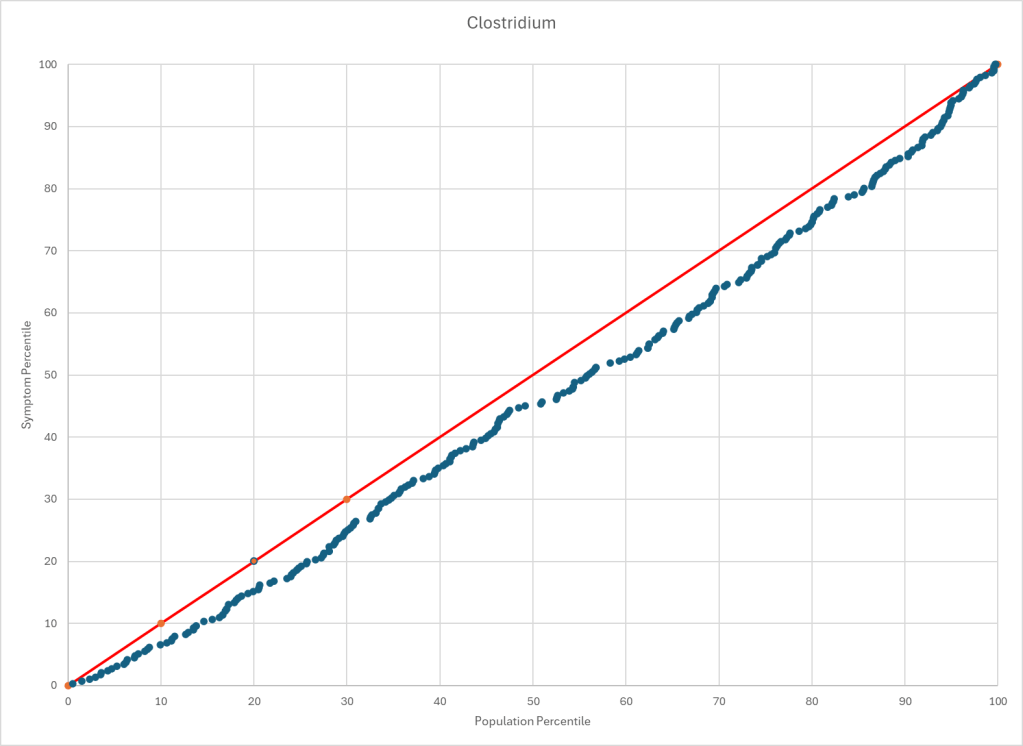
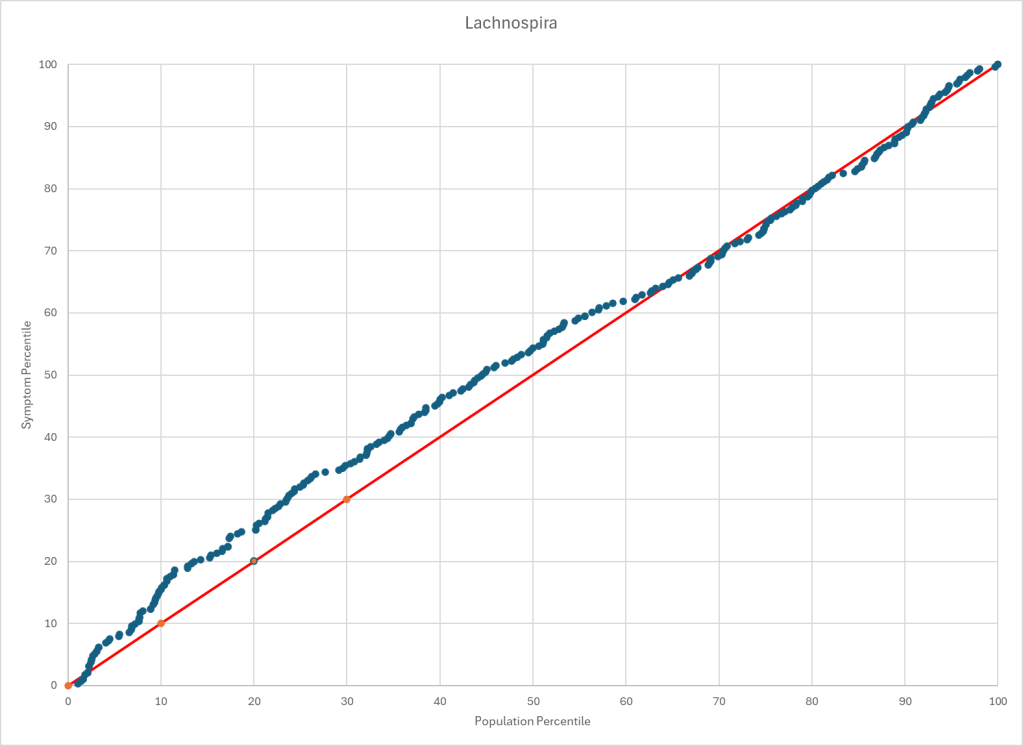
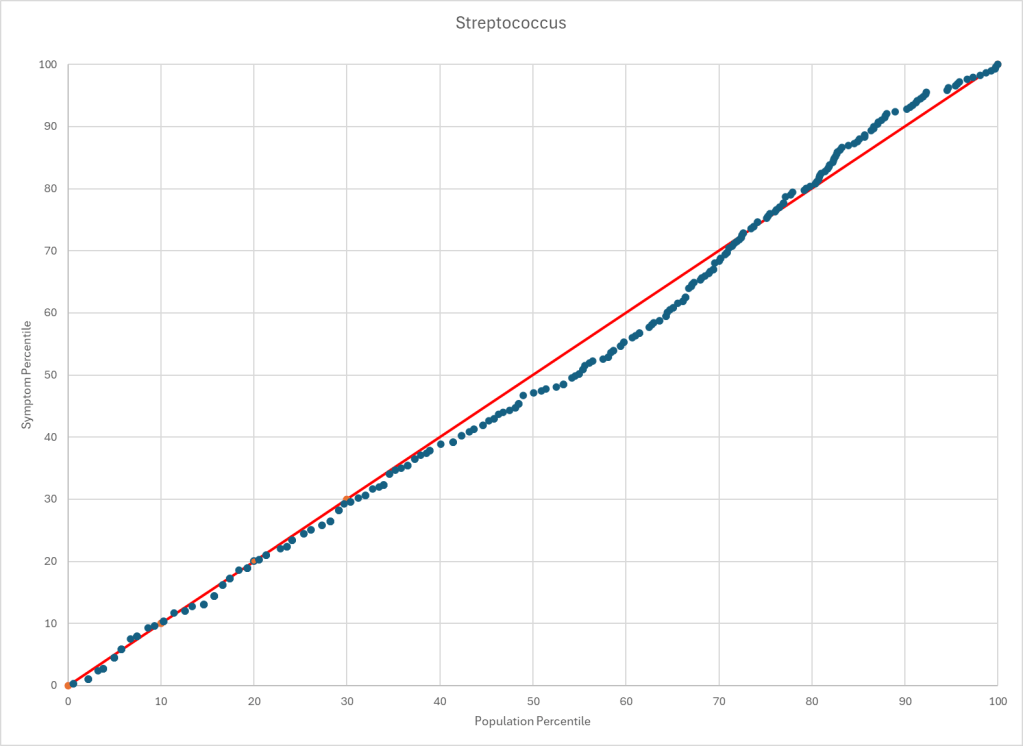
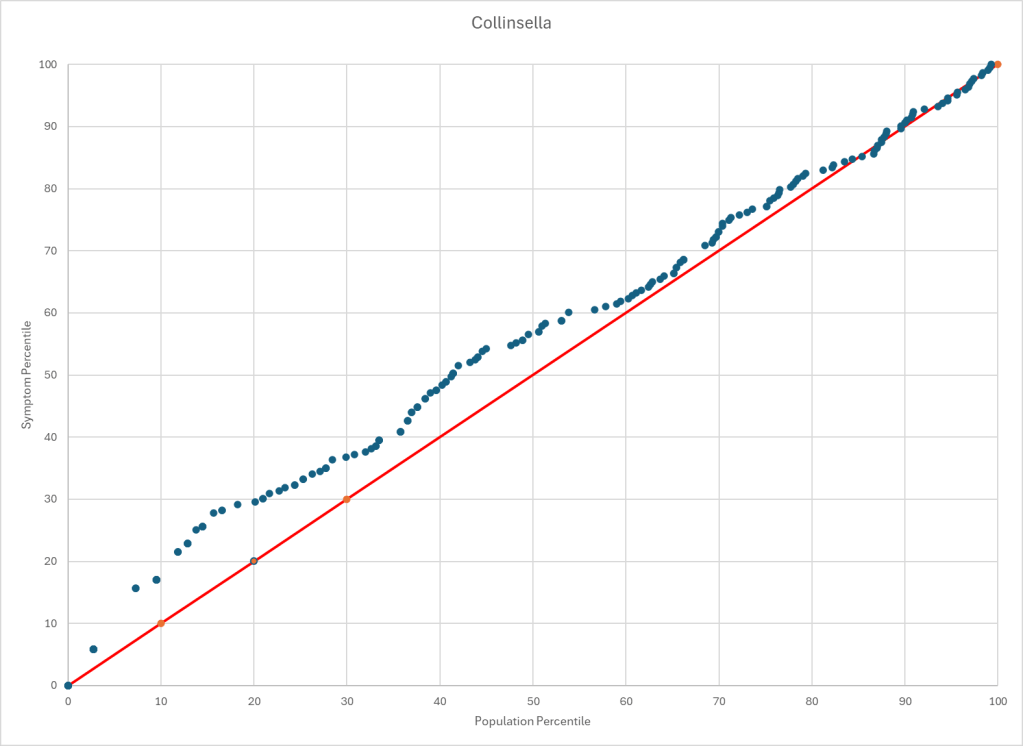
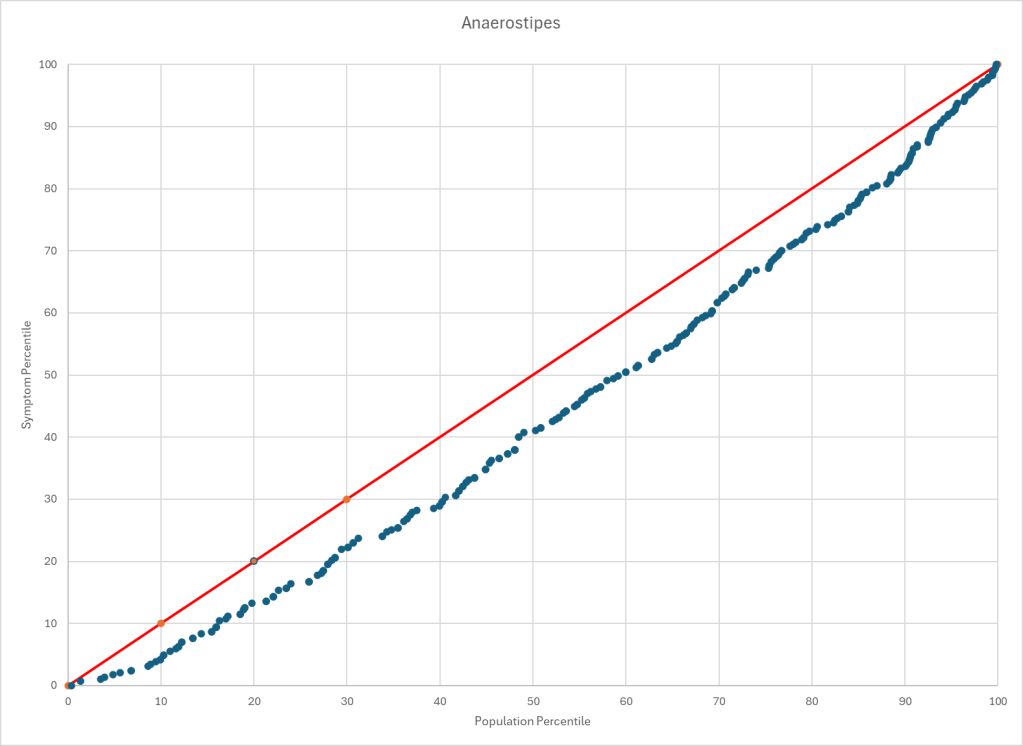
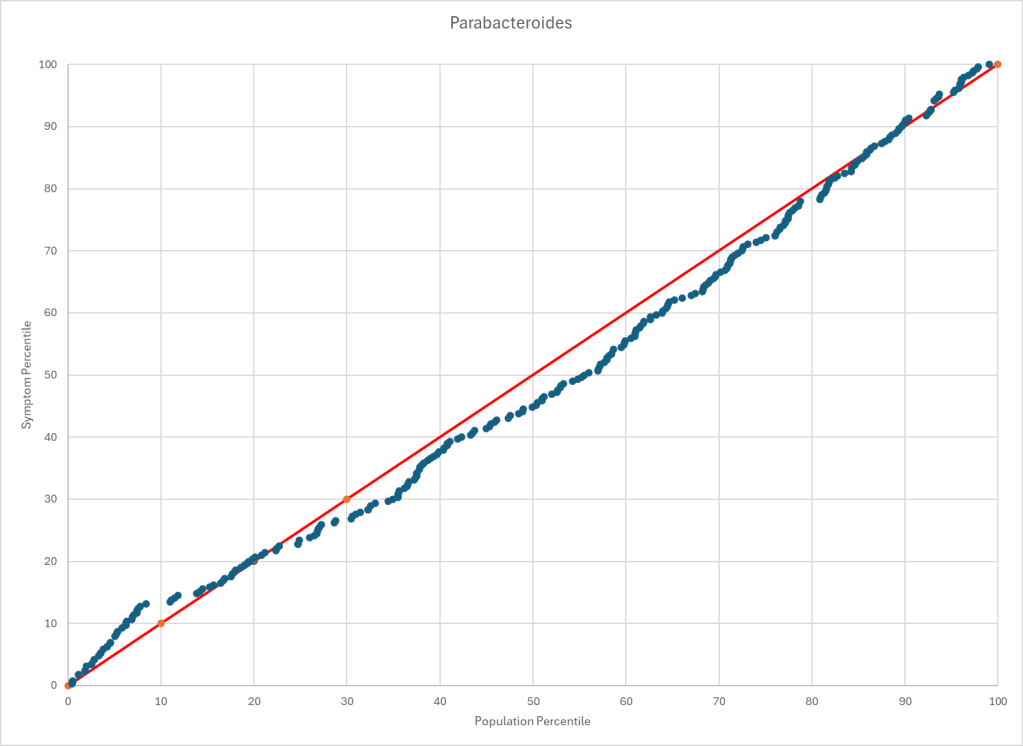
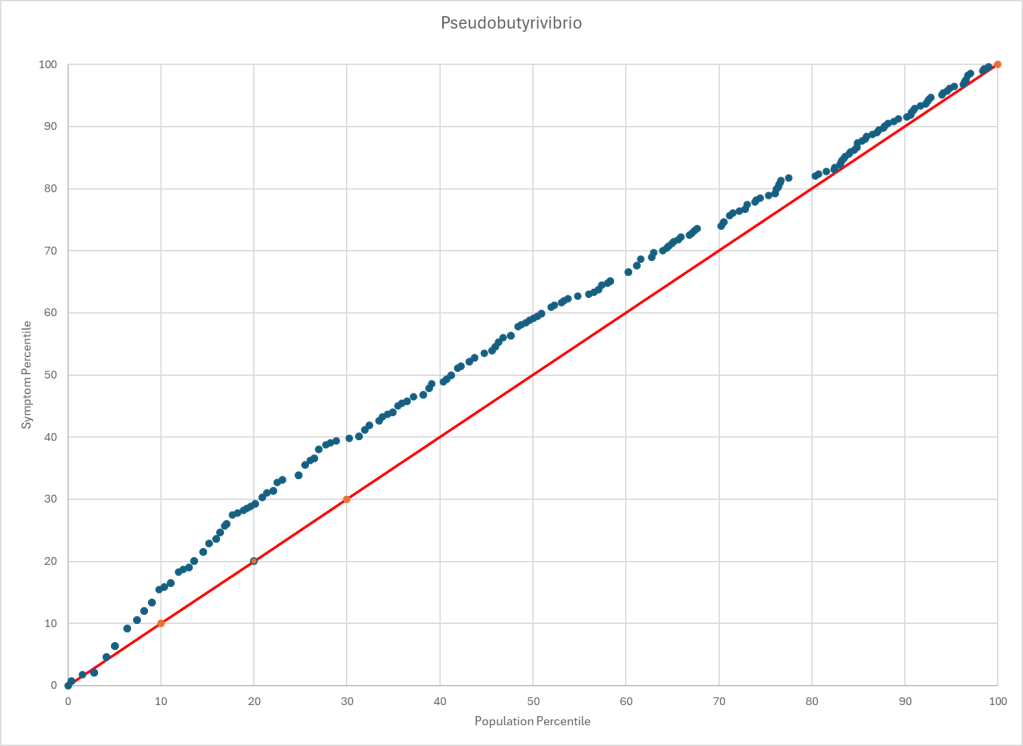
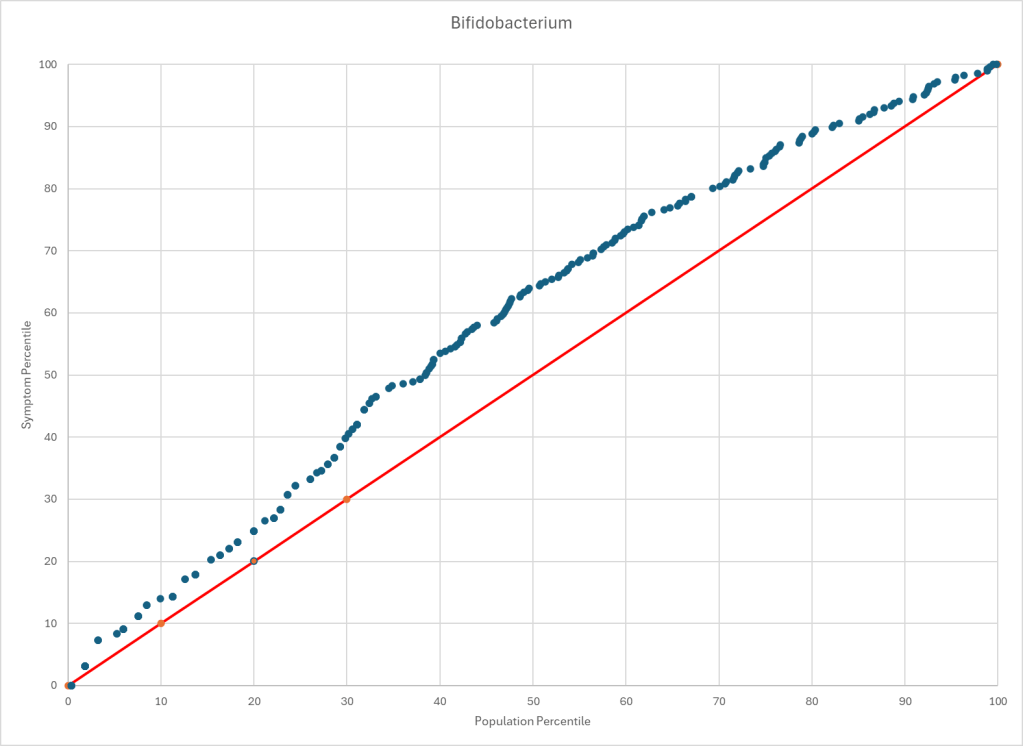
Statistics is fun because there many paths. Most studies using the microbiome uses the easy, but naïve, path of computing averages and standard deviation. As my dataset has grown, I have been travelling some less traveled path, for example: Visual Exploration of Odds Ratios, and a patent pending method termed “Kaltoft-Moltrup”.
One of the frequent decisions that I see in studies is to limit examination of bacteria that have a high frequency in the samples. This allows the researchers to keep to familiar and classic statistics. Using frequency of observation in the control group and the condition group is one of these much less travelled paths. It usually require big sample sizes and many studies have a sample size of 30 (sufficient for the mean and standard deviation approach).
I just completed code to compute Chi2 using Biomesight data for users reporting ME/CFS.
- Total Population: 3525
- ME/CFS: 280
Chi2 can be converted to probability (p) of happening at random with the following table

Seen too Rarely(Want to increase)
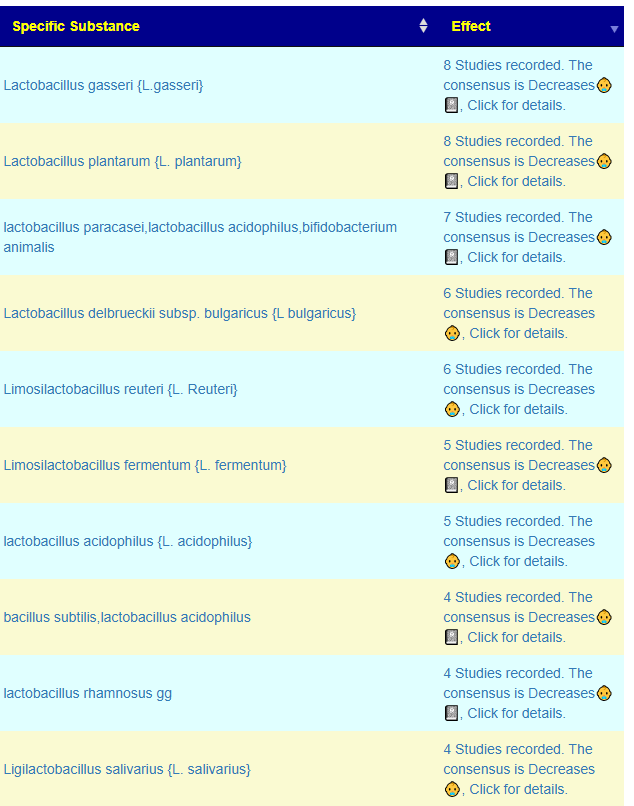
We see one bacteria available as a probiotic Bifidobacterium adolescentis. The rest would need to be altered by diet.
| tax_name | TAX RANK | Chi2 | Observed | Expected | Shift |
| Segatella oulorum | species | 13.9 | 16 | 38 | Under-Represented |
| Bifidobacterium cuniculi | species | 10.8 | 33 | 57 | Under-Represented |
| Brenneria | genus | 10.5 | 1 | 12 | Under-Represented |
| Bifidobacterium adolescentis | strain | 10.3 | 50 | 77 | Under-Represented |
| Prevotella veroralis | species | 10.3 | 6 | 20 | Under-Represented |
| Aggregatibacter | genus | 10.1 | 46 | 72 | Under-Represented |
| Hoylesella shahii | species | 10.1 | 12 | 28 | Under-Represented |
| Segatella paludivivens | species | 9.6 | 40 | 64 | Under-Represented |
| Actinobacillus pleuropneumoniae | species | 8.6 | 33 | 54 | Under-Represented |
| Slackia heliotrinireducens | species | 8.2 | 8 | 20 | Under-Represented |
| Prevotella micans | species | 8.1 | 3 | 13 | Under-Represented |
| Actinobacillus | genus | 8 | 123 | 157 | Under-Represented |
| [Actinobacillus] rossii | species | 7.1 | 79 | 105 | Under-Represented |
| Pasteurellaceae incertae sedis | no rank | 7.1 | 79 | 105 | Under-Represented |
| [Pasteurella] aerogenes-[Pasteurella] mairii-[Actinobacillus] rossii complex | species group | 7.1 | 79 | 105 | Under-Represented |
| Aggregatibacter aphrophilus | species | 7 | 4 | 13 | Under-Represented |
| Mitsuokella multacida | species | 6.8 | 3 | 12 | Under-Represented |
Seen too Often (Want to decrease)
We see 118 bacteria over a Chi2 of 6.635 ( P < 0.01 or 1 change in 100 of being a false detection). The list below are for genus and higher taxonomy orders.
| Tax Name | Rank | Chi2 | Observed | Expected | Shift |
| Dethiobacteraceae | family | 20.9 | 75 | 45 | Over-Represented |
| Hungateiclostridiaceae | family | 16.1 | 71 | 45 | Over-Represented |
| Tepidimicrobiaceae | family | 14.6 | 16 | 7 | Over-Represented |
| unclassified Burkholderiales | family | 12.6 | 29 | 16 | Over-Represented |
| Corynebacteriaceae | family | 11.2 | 159 | 123 | Over-Represented |
| Halanaerobiaceae | family | 10.1 | 76 | 54 | Over-Represented |
| Alcanivoracaceae | family | 9.6 | 53 | 35 | Over-Represented |
| Halanaerobiales | order | 9.6 | 81 | 58 | Over-Represented |
| Desulfonatronovibrionaceae | family | 9.1 | 58 | 40 | Over-Represented |
| Acetobacteraceae | family | 8.7 | 124 | 96 | Over-Represented |
| Hyphomonadaceae | family | 8 | 29 | 18 | Over-Represented |
| Tissierellaceae | family | 7.5 | 20 | 11 | Over-Represented |
| Rhizobiaceae | family | 7.1 | 98 | 76 | Over-Represented |
| Nannocystineae | suborder | 6.9 | 36 | 24 | Over-Represented |
| Rubritaleaceae | family | 6.9 | 124 | 99 | Over-Represented |
Bottom Line
The next step is to compute similar tables for all symptoms and incorporate these findings into a new algorithm. I say new, because I do not know if it is better than the existing ones. Conceptually, it would be added as a 5th set of suggestions to the existing consensus view on Microbiome Prescription.