Recently I have seen advertisements for smart watches for ME/CFS. The price would be challenging for many people. I suspect that it also has a poor return for the costs.
This post is about the watch family that my wife and I have been using for a few years. The current version is below. On occasion, I have seen the price under $20.00

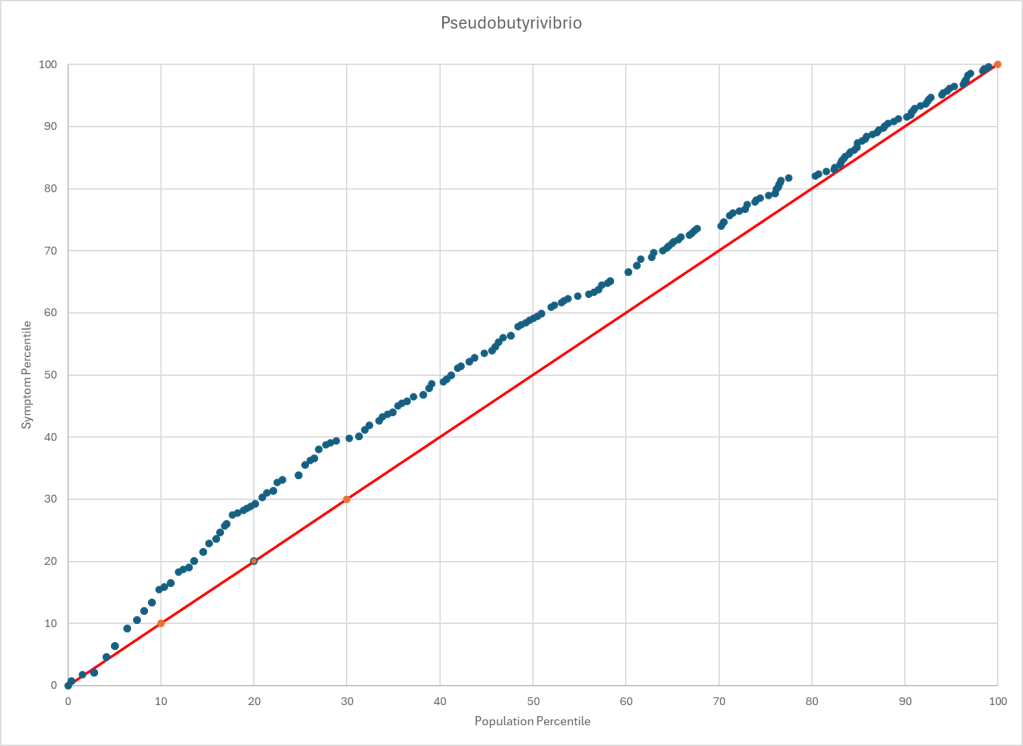
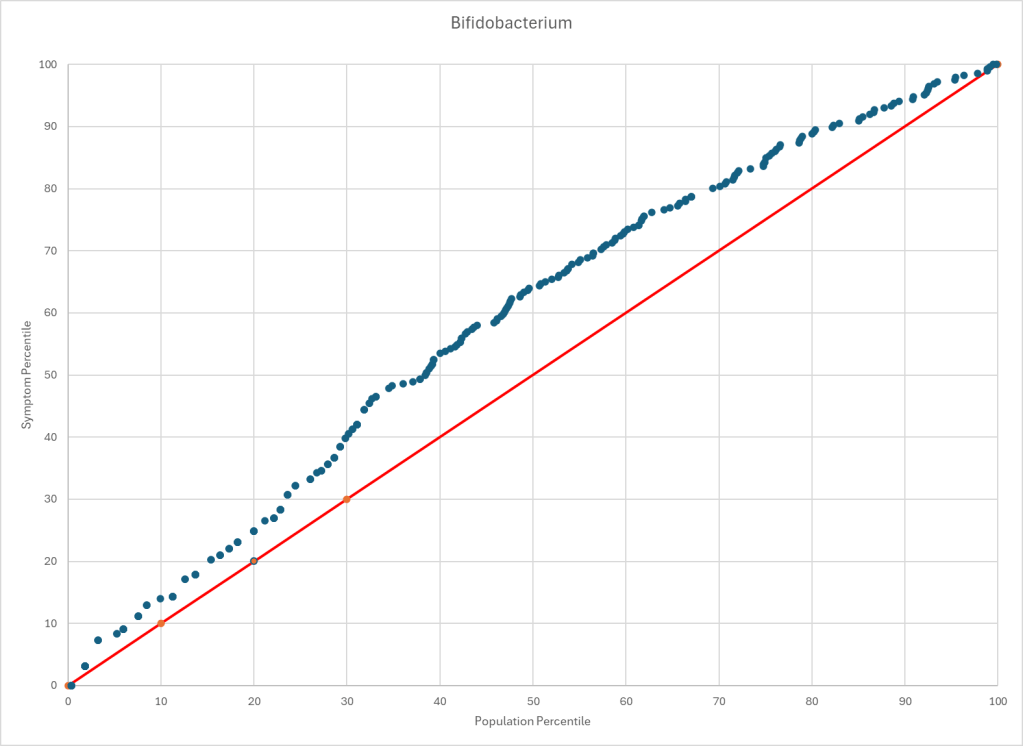
People who know me, knows that I prefer to work off objective data – for example microbiome test results, or actual physical measurements. This watch has empowered me. It can do automatic measurements every 10 minutes for day long monitoring.
What do I get with this watch?
You get an overview of the many measurements on a dashboard on your smart phone.
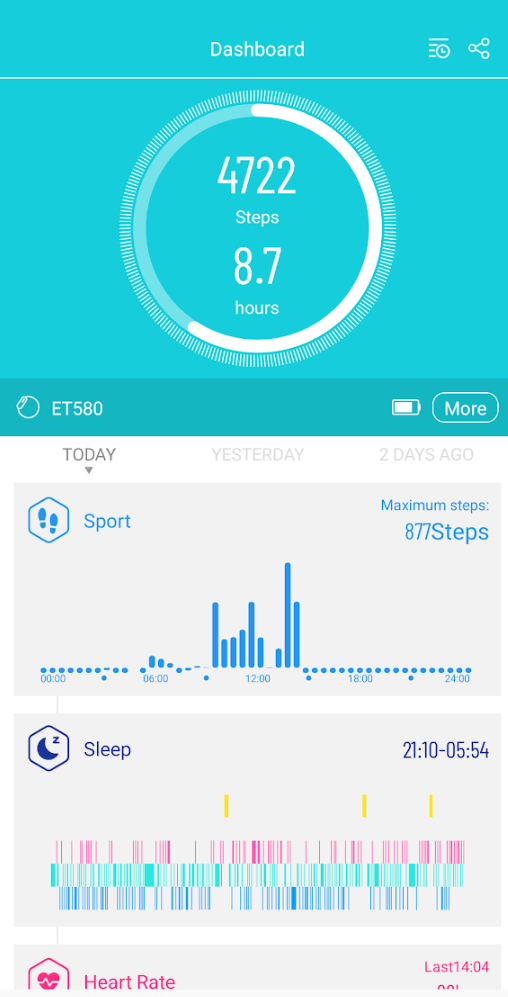
Steps per day

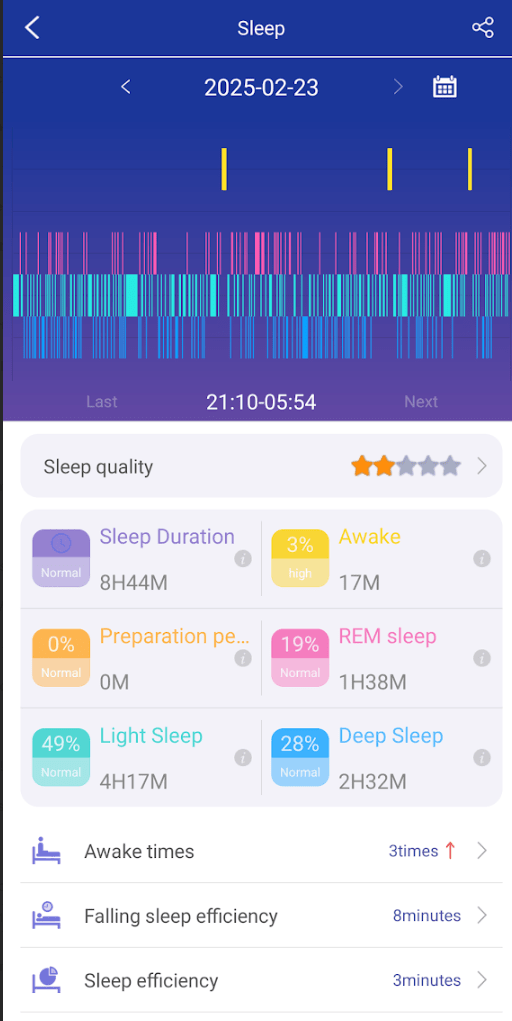
Sleep
Sleep is often a challenge. My wife found that some supplements improves her sleep and others do not. This changes decision making from being subjective to objective.
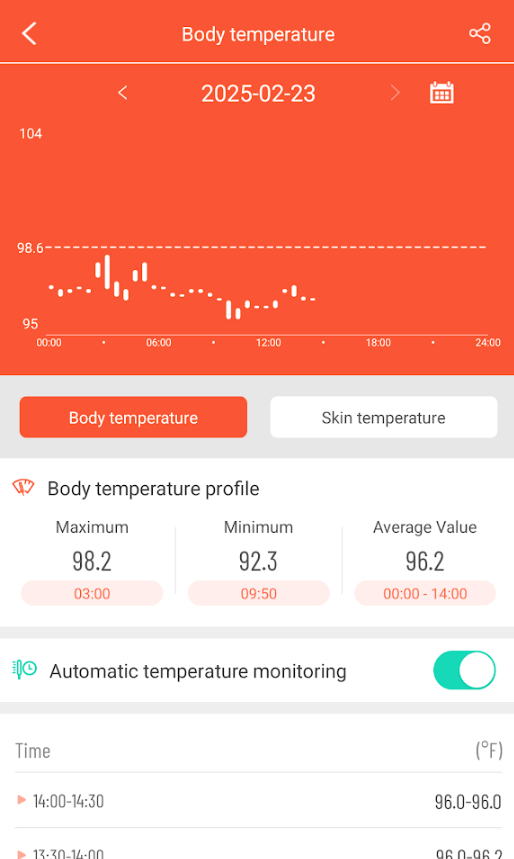
Body Temperature
As is common with ME/CFS people in general, I continue to have sub-normal temperatures. I notice that there is often a temperature spike while I sleep.
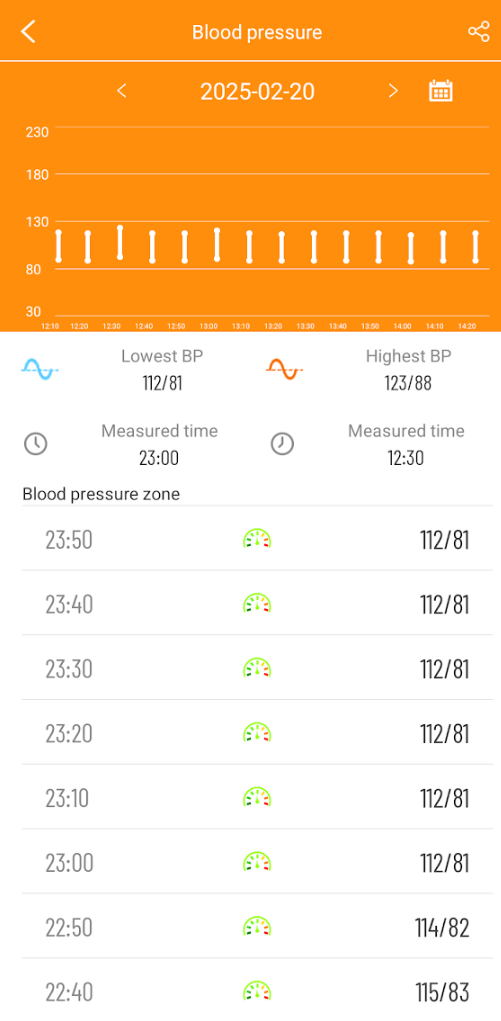
Blood Pressure
As often happens with ME/CFS, there can be low blood pressure as well as POTS.
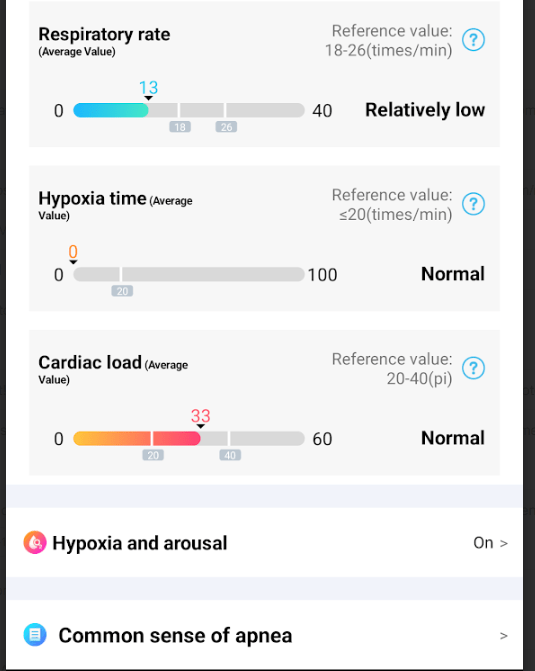
Blood Oxygen
My variant of ME/CFS had coagulation being a factor. This can result in hypoxia (low oxygen levels) causing cognitive issues.

Blood Glucose

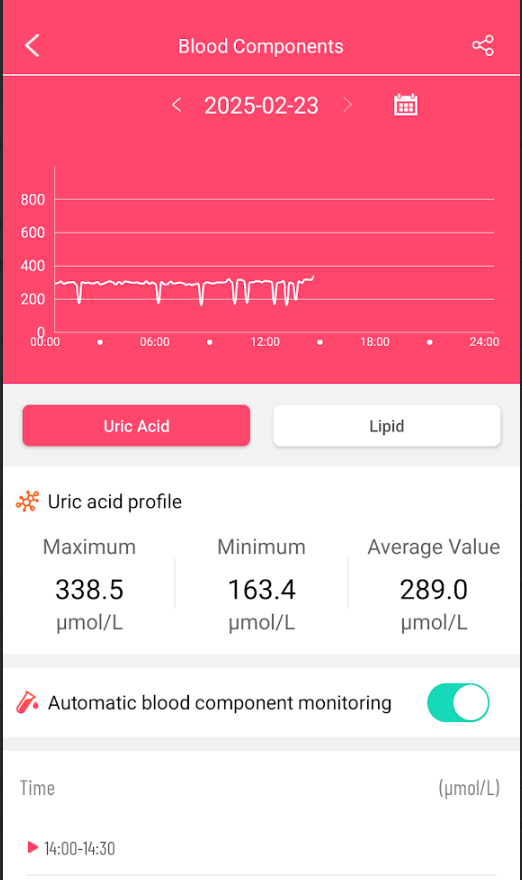
Blood Components

Heart Rate

HRV
Heart rate variability (HRV) is a measure of how much time varies between heartbeats. It’s a physiological marker of heart health and the autonomic nervous system (ANS). HRV can be used to assess physical and mental health, and to help diagnose diseases.
How HRV is measured
- HRV is measured in milliseconds
- A device is needed to measure the variation in time between heartbeats
- Wearable devices can be used to monitor HRV
What HRV indicates
- A high HRV indicates a healthy heart that can respond quickly to changes in the body
- A low HRV is associated with an increased risk of disease and death
- HRV can indicate activation of the parasympathetic nervous system (PSNS), which is associated with relaxation and good health
- HRV can indicate activation of the sympathetic nervous system (SNS), which is associated with stress and ill health

ECG Details
Heart Issues are frequently seen with ME/CFS. See The Heart and Blood of the CFS Patient


Overall Report

For example, I need to drink more water, etc.
Bottom Line
For less than $30, you get a lot of good information to act upon.
Questions
- Are the measurements accurate? In terms of being absolutely accurate — no. They appear to accurate for relative measurements: for example blood pressure. Items like skin color are known to impact accuracy of most optical sensors.
- How can it measure glucose? In terms of APPROVED FDA devices. There are no such devises for sale in the US. It is made in China (using the same type of sensors used in Apple, Google, FitBit, Samsung watches) and directly imported into the US “for personal use”, thus no FDA approval is needed. The FDA approval process is skipped and features available sooner.
- As always “buyer beware” — test it against “approved clinical grade devices” for measurements of significant concern. I have found that it seems very accurate for relative changes.