I will not look at the differences of what is reported. Many people have done blogs and articles complaining about differences. What I care about is the differences between analysis and suggestions — the actual application of the reports. Reader reports: “same sample, same time.”
Comparison tools
Reader Beware: There is no gold standard for matching rDNA to bacteria.- firms use different algorithms. Second, analysis is done using box-plot. The base data for box-plat is at present: 731 ubiome, 44 Thryve so there is a bias to ubiome’s processing.
1-15 is the Thryve sample, 7-11 is the uBiome sample

Thrye had more condition matches (113) compared to ubiome (88) – but is more bias to the low levels.
This may be explained by the number of bacteria reported, a major difference. The number for ubiome seemed very low (bad sample taken or processing issue).
- ubiome: 335
- thryve: 657
Let us look at what is reported by each.
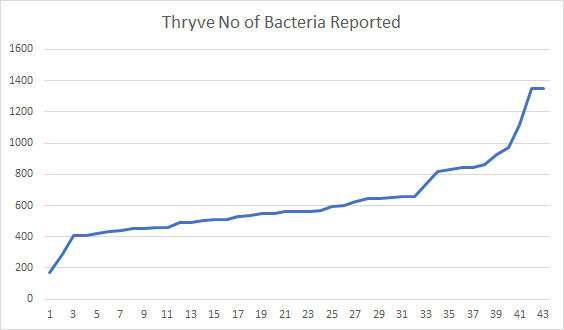
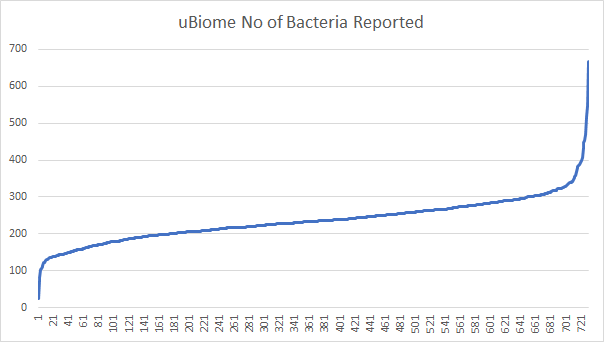
So we are at the 72%ile for Thryve samples and 96%ile for uBiome samples. The median counts: 234 for ubiome, 560 for thryve.
Conclusion: Thryve on average provide more bacteria reported. I look forward to comparing SunGenomics.com tests which should have even a higher count than Thryve.
Details
Looking at the highest level, Phylum, we see very great differences, especially in the more uncommon phylum.
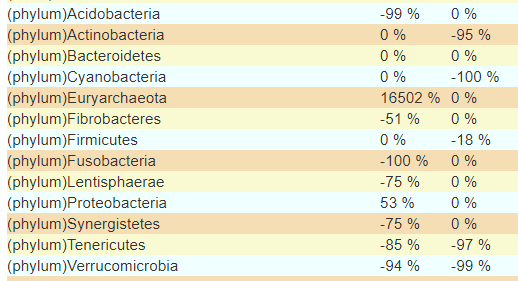
Suggestions
This is what most interests me. The suggestions prediction algorithms are not based on any lab results but on pubmed studies. The data used may be lab-dependent.
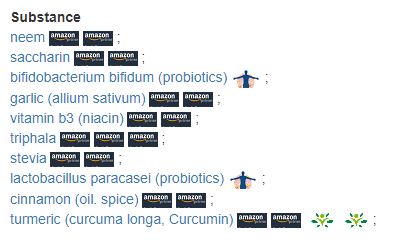

We have just one commonality: niacin.

Bottom Line
This has been interesting. Which is more accurate? I do not know; clearly results are very different (as reported by many other bloggers). Is the difference because of the computer algorithm used by each lab (known to be different), natural variation in a stool sample, sample collection process, etc.
Although the literature is generally conflicting with regard to sampling methodology, it is important to consider that comparisons of data obtained using different approaches should be avoided.
The Madness of Microbiome: Attempting To Find Consensus “Best Practice” for 16S Microbiome Studies
All steps of the complex analysis workflow significantly influenced microbiome profiles, but the magnitude of variation caused by PCR primers for 16S rDNA amplification was clearly the largest. In order to advance microbiome research to a more standardized and routine medical diagnostic procedure, it is essential to establish uniform standard operating procedures throughout laboratories and to initiate regular proficiency testing.
Multicenter quality assessment of 16S ribosomal DNA-sequencing for microbiome analyses reveals high inter-center variability
Before everyone goes to Thryve — be aware of the following:
- Box-Plot detection is based largely on ubiome samples. When I get 100 (at present 44, half-way there), I will split the reference numbers into a Thryve and a uBiome set.
- Averages were all reversed engineered from ubiome values.
- Symptom matching is based on submitted samples (94% from ubiome)
- Condition Templates are based on published research which uses a wide variety of tests methodologies.
So, the reality is that the site favors uBiome data.
There is a natural desire to get absolute, precise information. We are dealing with methods that are evolving. My attitude is to take the best data currently available (factoring in costs) and use it. It is better than working from no information.
