A reader asked me to write up a description for their MD. The PDF version is available here. Below is a text version
The difference between older Gut Labs and 16s Labs
Ken Lassesen, B.Sc. , M.Sc .
Interest in the microbiome has grown greatly in the last few years. Two decades ago, there was rarely 5 articles a month. Today it is close to 40 articles a day. A new laboratory technique called 16s has contributed to this explosion of studies.
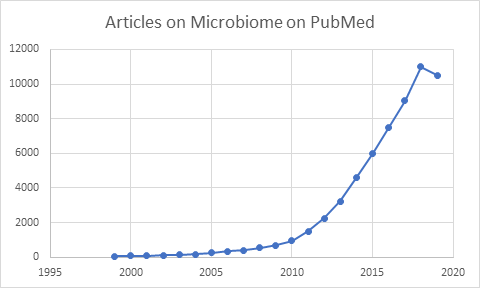
Practicing physicians finished their education before 16s usage became common. People like the familiar, and will tend to use tests that they have a long history with. A test like Genova Diagnostic Comprehensive Digestive Stool Analysis, has been in use since the early 2000’s with some changes over the years .
Methodologies
• Older Tests: utilized the standard culture method
o Many bacteria cannot be cultured, in fact most cannot be.
• “The 16S rRNA gene is a short section (~1500 base pairs) of prokaryotic DNA found in all bacteria and archaea. The 16S rRNA gene is a commonly used tool for identifying bacteria because analysis of an organism’s DNA is often more definitive than classification based solely on phenotypic characteristics.”
Translation:
• Older Tests often report on 20-30 bacteria only
• 16S Tests typically report on 3000-4000 bacteria
Distributions and Determining Outliers
The traditional labs follow the tradition lab process to set limits of what is normal. “A minimum sample population (20-25) samples is all that is required for transference of reference ranges.”
From this number, they compute an average and a standard deviation and then commit a fatal flaw in dealing with the microbiome. They assume that they have a normal or bell curve distribution.
This same naïve mistake is often repeated at laboratories offering 16s services.
To illustrate this, we have 146 samples with Lactobacillus crispatus with an average of 7800 but 90% of the samples were less than 7800. The plots below show values of different taxonomy
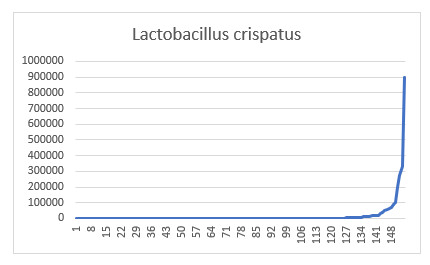
On the other hand Bacteroidia Class is better behaved, but with mean +/- 2 standard deviation
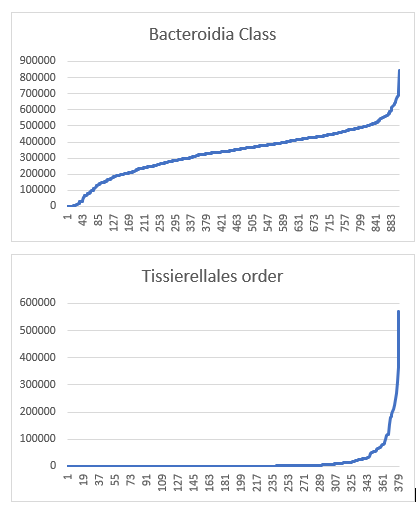
The Correction Solution
The correction solution is to compute the actual distribution bacteria by bacteria with a sufficiently large sample and then determine what is in the top or bottom 1%, 3% or 5% of reading.
What is your belief?
Which test is best, partially depends on your belief as to the cause of the problem.
• The traditional (simple and familiar) test assumes one or two bacteria are the cause and from a small collection of candidates.
• The 16s view of a microbiome dysfunction is more complex with interactions between bacteria supporting or inhibiting each other.
•
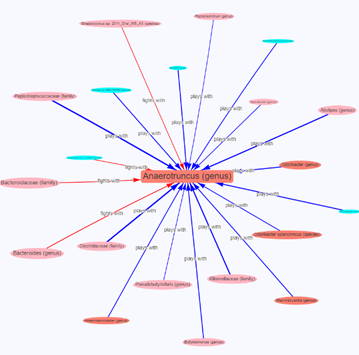
The more complex view with perhaps a hundred times more bacteria is a challenge. One solution to this is the use of artificial intelligence techniques.
Missing Bacteria (or apparently missing)
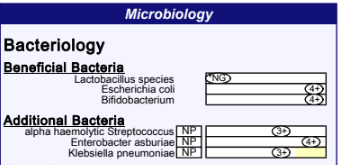
The biggest challenge when comparing reports is that 16s does not have a 100% match to the traditional reports. The most common complaint is that Escherichia coli appears missing. This is a case of “rose by another name” combined with a little politics over the boogie man called ‘E.Coli’ which is traditionally viewed by most retail consumers as unhealthy and dangerous.
Kluyvera and Escherichia genus are not distinguishable using the common 16s rna primers. The primers work only on a portion of the complete DNA and it (unfortunately) happens that this portion have strong similarity between these two. That combined by the bad press E.Coli gets, has likely caused retail testing companies to go with the little known bacteria Kluyera instead.
Bottom Line – Which is Better?
This really depends on the model that you are working with:
• It is dues to just one or two bacteria and you want a simple view without having to deal with interactions and complexities – then the traditional tests is best.
• If there are multiple bacteria involved (and most of the research appears to confirm that is the case for many diseases including CFS, Parkinson’s, Alzheimer’s, Diabetes) then you want to use the 16s test and hopefully have access to appropriate tools to evaluate it because there will be massive information overload.
Personally, I believe 16s is the better of the two – a lot more information (and thus more effort needed).
Units of Measurements Are Different
Most 16s reports are units per 1,000,000 bacteria (often the actual count is less, often just 10%, i.e. 100,000 bacteria).
count
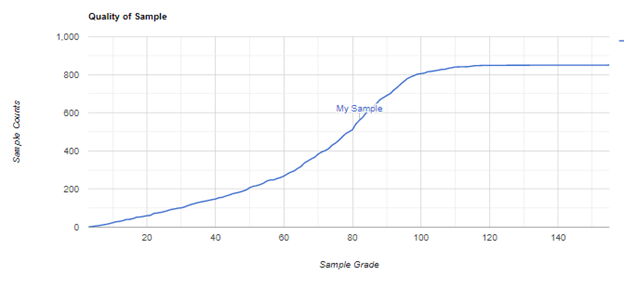
Figure 2 From 800 samples Counts are in 1000’s
The traditional lab gives a count in Community Forming Units/gram. The use of CFU/g clearly states that the method used is culturing. The size of bacteria seen in nature is reported to vary by 10 orders of magnitude . Weight (as in grams) is the cube of the size dimension, so the weight will vary by 1000 orders of magnitude. Culturing also puts a bias in the relative numbers because some culture better than others. Looking at one lab report, I see reference ranges of 1.5E10 for Roseburia spp and 2.4E5 for Fusobacterium – a difference of 100,000x (E5). Looking at the 16s data from 800+ samples, the median numbers are 368x difference, and the means are 6.4x different.
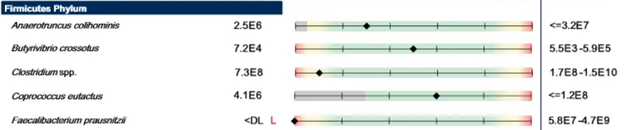
Some labs use their own scales which makes it impossible to correlate with other reports is a meaningful way.

Current thinking
The ‘pathobiome’ — a new understanding of disease, University of Exeter, Sep 12, 2019 https://www.sciencedaily.com/releases/2019/09/190912113238.htm
“Traditional approaches to describe infectious disease in plants, animals and in humans are based on the concept that single pathogens are responsible for the signs or symptoms of disease observed in those hosts.
The concept acknowledges that all organisms are in fact complex communities of viruses, microbes and other small organisms (e.g. parasites) which can interact to affect health or disease status at any given time.
These complex communities continually interact with their hosts, sometimes conferring benefits (e.g. “good” bacteria in the human gut microbiome), and at other times causing harm by contributing to disease.
When these communities combine to cause disease, they are termed “pathobiomes” — a recognition of their collective shift away from the healthy-state “symbiome.”
The recognition that the pathobiome plays a key role in those signs and symptoms of disease that we observe in the host is becoming a more accurate way of considering disease than by simply referring to it as the outcome of the effects of a single pathogen (e.g. the influenza virus).”
At the citizen science site, http://microbiomeprescription.com/, symptoms (and not diagnosed disease) were found to have very strong correlation using Pearson’s chi-square tests.– even for some age ranges,

This may explain why many studies have contradictory results. The condition group has a diagnosis, for example Chronic Fatigue Syndrome which is defined as:
“The Centers for Disease Control and Prevention criteria for chronic fatigue syndrome include severe fatigue lasting longer than six months, as well as presence of at least four of the following physical symptoms:”
If symptoms are tied to bacteria shifts, then mixing people with different symptom sets (but the same diagnosis) generates a lot of statistical noise.
Bottom Line
Using 16s tests opens the door to treat for the symptoms and not for the diagnosis. This also opens the door for uber-personal medicine based on the person’s microbiome. Two people with the same diagnosis (and perhaps significant symptom overlap) may be put on very different treatment plans.
A simple example: suppose two persons are suffering from d-lactic acidosis. The bacteria producing d-lactic acid may be very different. One person may have overgrowth of lactobacillus Another person may have Klebsiella pneumoniae and Candida freundii overgrowth .
