This is an analysis of a parent with two children with a mixture of challenges. A person’s microbiome and their DNA appears to be related. Ignoring studies, it makes logical sense. The DNA favors certain metabolites, which favors certain bacteria — and the reverse.
Perturbations in the gut microbiota have been linked to atopic diseases. However, the development of atopic diseases depends not only on environmental factors (like microbial stimulation) but also on genetic factors.
Host-microbial interactions in childhood atopy: toll-like receptor 4 (TLR4), CD14, and fecal Escherichia coli [2010]
The family:
- P – Parent: No ASD, nor CFS. No current IBS (I had IBS symptoms 2003-2012). Sample: BiomeSight
- A : ASD/CFS Sample: Thryve, FASTQ processed thru BiomeSight
- B : No IBS. He had SIBO in 2017. Has been in remission from PANS for about 1 year prior to sample. Sample: BiomeSight
They live in the same household with similar meals.
Looking for Family Shifts
I looked first for bacteria that were high (top 6%) or low (bottom 6%) across multiple samples and found some interesting patterns:
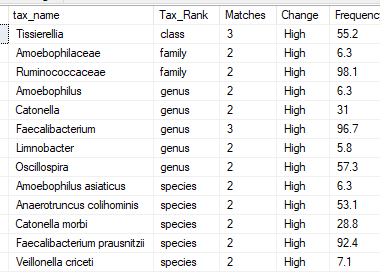
Items like Limnobacter and Amoebophilus really jumps out. They are rare, so the odds of 2 people at random having them and both being in the top 6% is about 0.002%. Adjusting for BiomeSight samples only, the odds decrease but still less than 1%.
Tissierellia and Faecalibacterium were high in all three samples.
The unfortunate aspect is that of these 4 bacteria, we only have significant data on how to reduce Faecalibacterium. Checking PubMed on Limbobacterm, we see barely 40 studies with the count being low in Parkinson’s disease [2018]
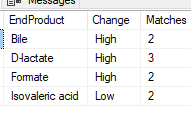
End Products Shift Shared
Applying the same criteria for End Products being produced, we have d-lactate being #1. This is common with CFS and I have written about it before (see posts below). It is also likely common with ASD.
- Reducing Lactic Acid Producing Bacteria – LABO
- Approaches to D-Lactic Acidosis
- Reminder of D-Lactic acidosis and ME/CFS
For which bacteria produces which end products, see this page. I have on my backlog of features a page that makes suggestions based on end products to be shifted only. At the moment, it’s a manual process of hand picking bacteria.
KEGG Module
Here, we will use top and bottom 3% since we have a lot more categories to examine. We found no shared items.
KEGG Enzymes
Here, we had only one hit.
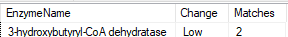
Bottom Line
The intent of this post was to look at commonality over a genetic related group of people living in the same house and likely eating the same diet.
For individual suggestions by person, I would advocate doing a hand picked taxonomy and working from there. For the child with ASD/CFS, I went and did that with their sample
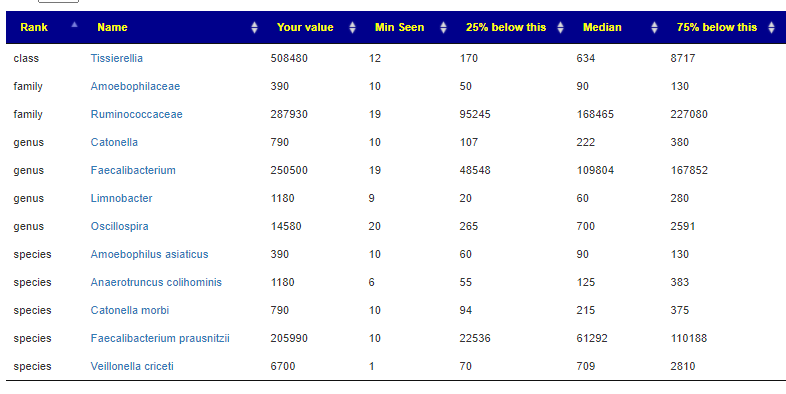
The resulting suggestions are shown below:
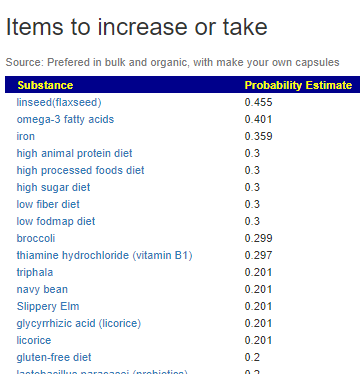
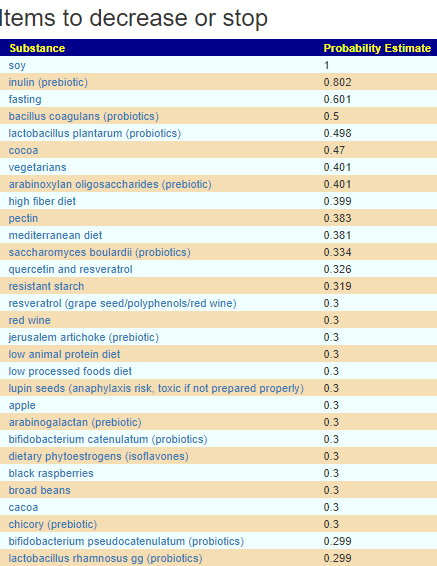
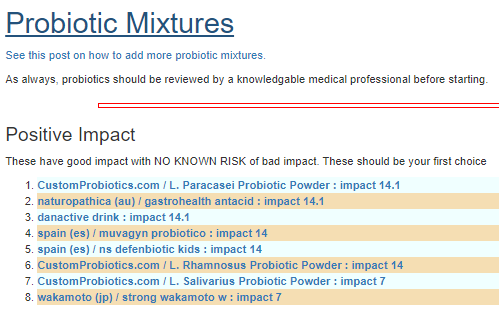
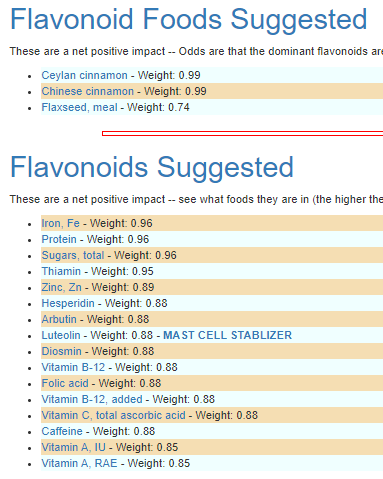
As always, these are suggestions that should be reviewed with your medical professional before doing. They are items believed to have better odds of making favorable changes than picking random items to try.
Future Pages
Most of the analysis above was done via ‘the backdoor’ and not available yet on the web site. The analysis code is written, it’s just a matter of getting time to create and test the pages.
