A reader has been kind and granted permissions to use their samples for another analysis. The samples were done by Thryve, and the FASTQ files processed thru Biome Sight. There was also a GI-MAP result available.
The medical diagnosis is Seizures Lennox-Gastaut syndrome to be exact. And severe low functioning autism. It is suspected/speculated that the cause was vaccine injury.
Enzyme Analysis
First I am going to look at the last sample and then do a comparison to the three prior ones. I am starting to feel that the Enzyme analysis is a good indicator of microbiome dysfunction. It measures thousands of enzymes using the genes found in the bacteria found.
- Thryve: Literally 139 Items, all HIGH VALUES
- BiomeSight: 131 items most high with 9 low and 12 low.
Probiotic suggests are only for LOW values (i.e. bacteria that produces the enzymes that you are low in). The list looks similar to prior analysis with lactobacillus paracasei being prominent.
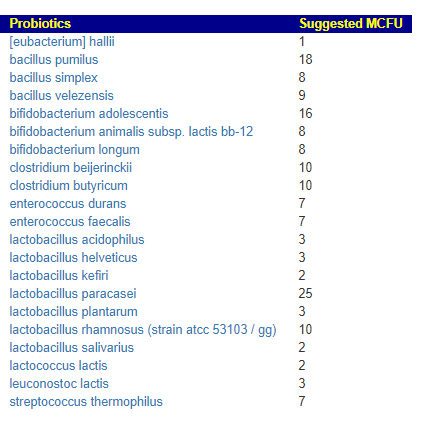
Comparing latest to prior, we found
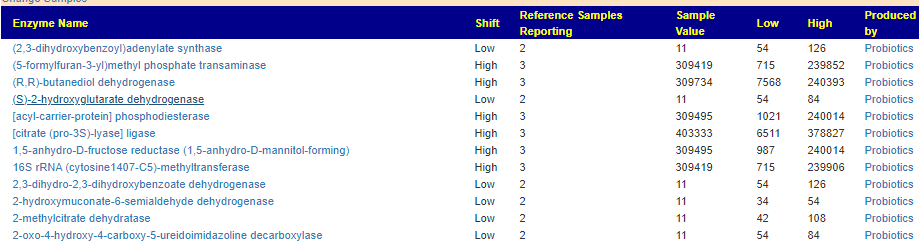
- Thryve: Literally 190 Items being outside of the earlier ranges. Some increasing and some decreasing, the general pattern is ones with high ranges are going higher, and those with low ranges are going lower 😦
- BiomeSight: 36 items all showing lower values than the 3 prior samples
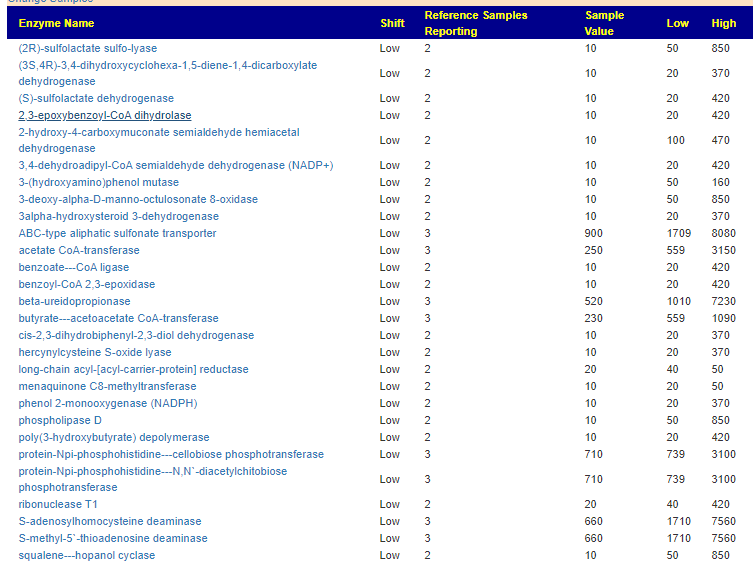
In short, there appears to be a major dysfunction for the enzymes being produced, and it seems to be getting worst.
But…. For End Products, with both samples, there was no change from prior samples. Thryve alone reported possible outliers — all of them them high values (thus no sense in taking supplements for these)
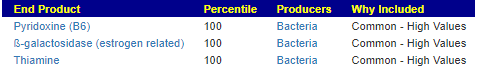
For Bacteria Changes:
- Thryve showed an major increase in Clostridium intestinale, Bacteroides uniformis, Bacteroides cellulosilyticus, Alistipes shahii, Mycoplasmatales with the values being high compared to other uploaded samples.
- BiomeSight showed major increase in Bacteroides uniformis, Bacteroides cellulosilyticus, with the values being high compared to other uploaded samples.
GI MAP Report
The following abnormalities were reported.
- Escherichia spp. – 50% below low bound – Not reported by Thryve or Biomesighe
- Akkermansia muciniphila – 4700% above high bound (between 30-65%ile on samples)
- Firmicutes – slightly above high bound (between 20-70%ile)
- Bacillus spp. – 7800% above high bound (all between 50-75%ile, except for one at 15%ile)
- Enterococcus faecalis – 8230% above high bound (none reported)
- Pseudomonas spp. – 132% above (all between 50-70%ile)
- Streptococcus spp. -2920% above (all between 28%-60%ile)
In short, none of the samples are in agreement with the report from GI-MAP. Personally, I do not have GI-MAP on my reliable/useful lab reports — the above differences are just too troubling.
Approach: Backtrack from Enzymes to Bacteria
These samples are a problem for traditional analysis. Individual bacteria may well be in the normal range, but may be in the high end of these ranges, causing too much enzymes to be produced when a bunch occur at the same time.
The result was my creating a novel analysis process: Compute by backtracking from the enzymes to the genes that produces the enzymes, and then use these genes to backtrack to the species and strains that have those enzymes. From both labs reading of FASTQ files, we get the same dominant players as shown below. Faecalibacterium prausnitzii was interesting — because of the high count, many will assume it to be a major player… it’s not (remember the counts for this bacteria will be different from each lab).
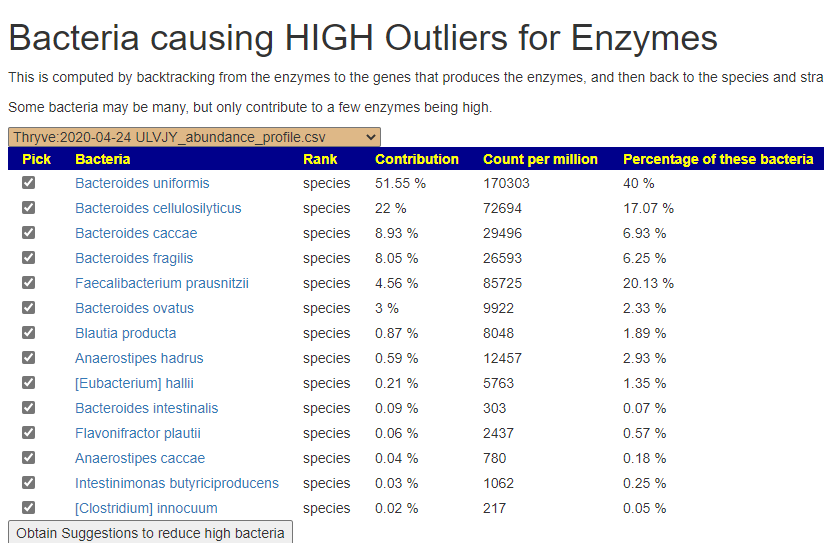
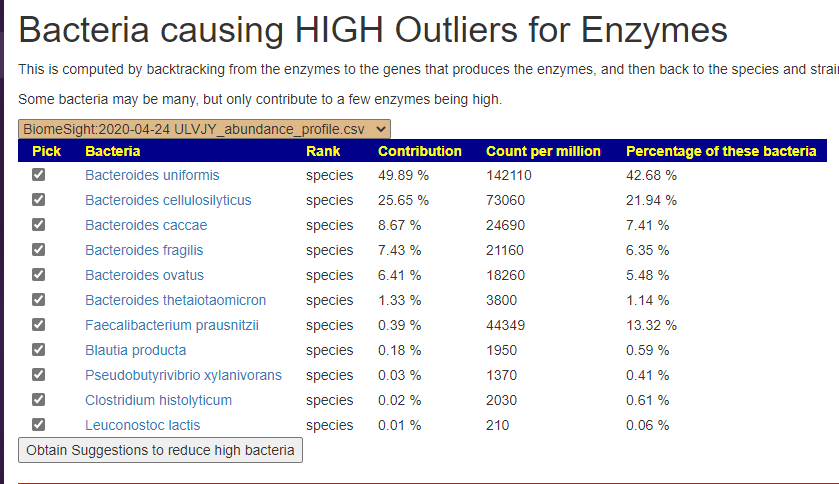
I then wired this into the existing suggestion system using the hand-picked bacteria approach (all are pre-selected on this page). You will now see a new button on the sample pages.
You can then custom tune the suggestions you wish to receive. For example, restricting to drugs and herbs, we have:
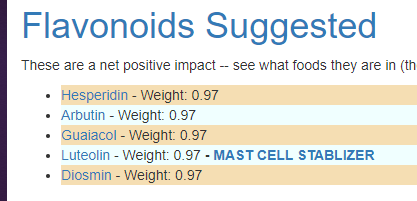


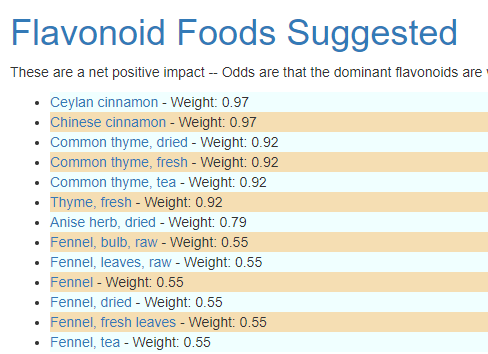

Pass 2
Since we are entirely at the species level, we may wish to include items that modify the genus these species are in. This is a simple change.

Suggestions were very similar without significant changes. Including probiotics, nothing showed a reasonable confidence (i.e. lactobacillus paracasei (probiotics) was a 0.038, I usually stop at 0.5)
Bottom Line
There are many paths for getting suggestions. For this person, the usual paths had been tried before with little success from experts that I know. When I saw the Enzyme madness, I realized that the first goal, a measurable goal in subsequent samples, is reducing these high enzyme levels.
To do this, I had to add more algorithms to the site to allow a backtrace from enzymes to bacteria.
In checking the literature for Epilepsy, I found the following articles which may be worth reviewing
- Gut Microbiota, the Ketogenic Diet and Epilepsy [2018] – no symptom improvement cited with KD diet
- Current Perspectives On The Role Of The Ketogenic Diet In Epilepsy Management [2019] “Multiple trials have demonstrated that KD is an effective treatment modality in children with Dravet Syndrome” but not for this person’s condition
Of interest/concern is that with KD diets, “after KD therapy and revealed significantly decreased abundance of Firmicutes and increased levels of Bacteroidetes.” In this case HIGH Bacteroidetes appear to be the issue, at least specific species, changing the Bacteroidetes species may result in positive results — speculation.
Now going to the other aspect, Autism
- Gut Microbiota Features in Young Children With Autism Spectrum Disorders [2018]
“22 Bacteroidetes OTUs owing to genus Bacteroides, mainly assigned to B. uniformis and B. vulgatus and P. distasonis species, were more abundant in ASD patients … an increase of Faecalibacterium prausnitzii, “ - Fecal Microbiota and Metabolome of Children with Autism and Pervasive Developmental Disorder Not Otherwise Specified [2013] Table 2 list many of the species that we identified above.
My conclusion, for these samples, is that looking at the enzyme shifts lead us back to the same bacteria already identified for autism. What we’ve added is that a contributing factor for autism may be the overproduction of specific enzymes which may lead to alternative treatment approaches. Using this as an assumption, we also can rank order the most important ones to change (i.e. Bacteroides uniformis – links to a list of items that inhibits this specific bacteria)
Recipe book to discussion with Medical Professional
- Vitamins:
- Vitamin C (ascorbic acid)
- B Vitamins ( B1, B3, B6, B7, B-9, B12)
- Melatonin supplement
- Herbs and Spices: Cinnamon, Thyme, Anise, Fennel
- Supplements
- Probiotics:
- CustomProbiotics.com / L. Casei Probiotic Powder :
- schiff / digestive advantage (any that contains bacillus coagulans – I believe there are some gummies available with it)
- Foods
- whole-grain barley (ideally as porridge every day)
- walnuts
- momordia charantia(bitter melon, karela, balsam pear, or bitter gourd) – may be an acquired taste
None of the following: Licorice, resistant starch , tannic acid (cranberries, strawberries, blueberries, apricots, barley, peaches, mint, basil, rosemary), stevia, saccharin
Toss up on apples — some of its ingredients may help or make worse. Do as an experiment when he is stable.
Remember this is not medical advice, before doing any changes make sure that you review them with your medical professional. This is what a data nerd logically deduced (showing their logic), and not a qualified medical professional.
