A reader from Romania requested that I do a post on MCAS. It’s of special interest to me because some close friends appear to have this issue. I know from prior posts that there is limited professional knowledge that is effective, and most of it is symptom relief and not addressing the underlying cause.
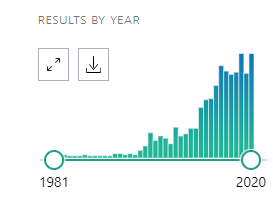
I recently added a major component to my microbiome analysis site – enzymes from Kyoto Encyclopedia of Genes and Genomes. Witg this, we have had a significant number of microbiome samples shared to the site reporting Comorbid: Histamine or Mast Cell issues. One of the challenges is that there may be some fuzz since this is self reported.
In this post you will see that enzyme imbalances from a microbiome sample can lead to a root cause of some symptoms. In some cases, lead to actions that may moderate the symptom.
The process that I going to use, is simple:
- Look at each species reported in each of 224 people microbiome (over 10,000 different bacteria species)
- Look up the genes in each one of these species.
- Look up which ones of 1600+ Enzymes each gene produced. Most bacteria produces many many enzymes – they have a lot of genes.
- Look at statistically significant oddities in the enzymes of those reporting a certain symptom
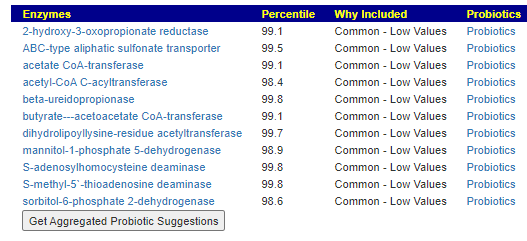
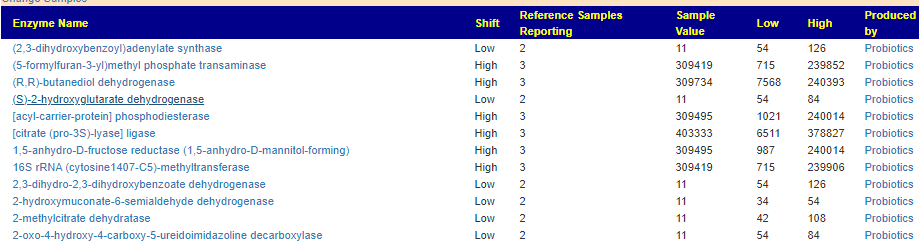
When this data is pushed thru some artificial intelligence, one enzymes leaps out for MCAS, lipopolysaccharide 3-alpha-galactosyltransferase.

Four other enzymes are also suspect (and low) but without as strong statistical significance at present, as the one above.
- aureolysin
- lysostaphin
- poly(ribitol-phosphate) alpha-N-acetylglucosaminyltransferase
- 4,4
-diapophytoene desaturase (4,4-diapolycopene-forming)
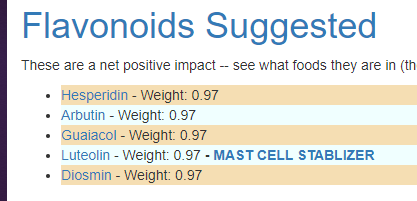
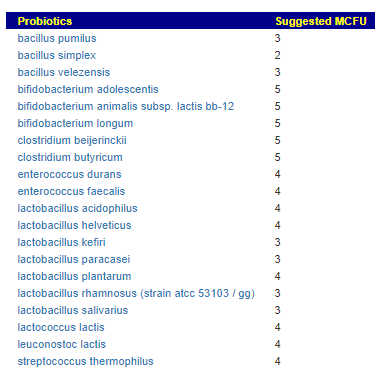
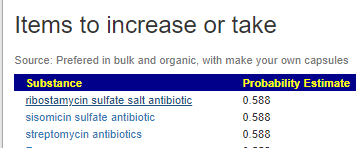
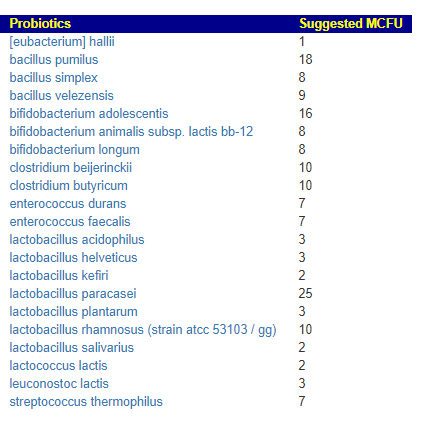
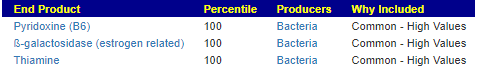
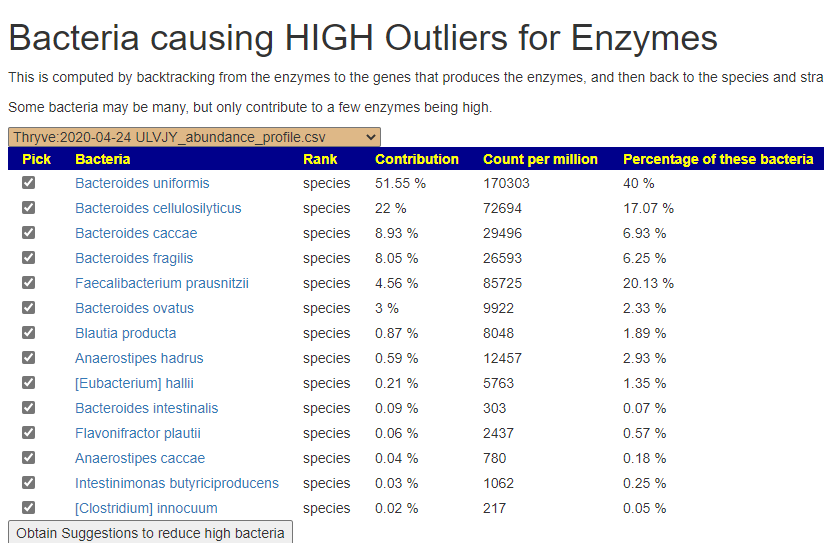
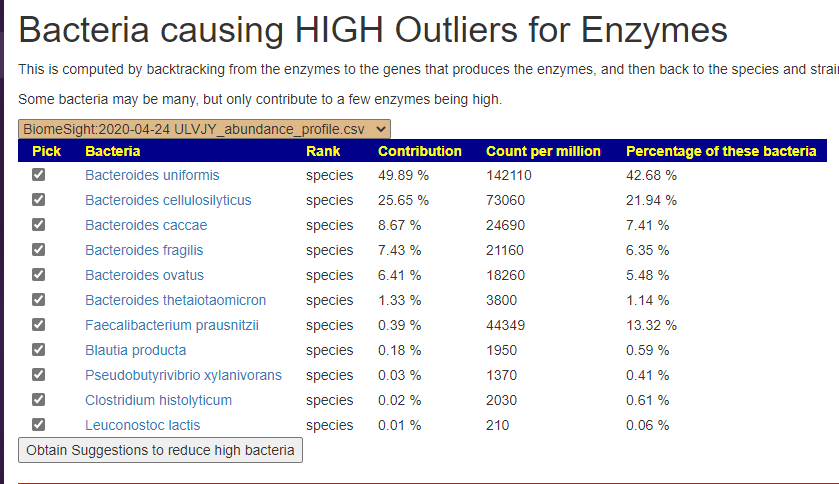
We can also walk this bacteria-to-enzyme path backwards, from the enzymes that are low, we look up the genes that produces it, then from the genes identify the bacteria; as a final step, we see if any commercially available probiotics produces this enzyme. The thinking is simple: if you are short of an enzymes, then increasing enzymes may calm down the processes causing MCAS. We come up with two of them:
This looks like AI magic, machine learning of other evils of technology. Not quite, we can look for studies dealing with allergies and/or MCAS and see if either of these have been studied. First, we discovered that is not a single study on using the above to treat MCAS. For allergies, histamine reactions, we find significant literature
- Lactobacillus helveticus SBT2171 Alleviates Perennial Allergic Rhinitis in Japanese Adults by Suppressing Eosinophils: A Randomized, Double-Blind, Placebo-Controlled Study [2020]
- Lactobacillus helveticus SBT2171 alleviates allergic symptoms in a murine model for pollen allergy [2019]
- Modulation effect of Lactobacillus acidophilus KLDS 1.0738 on gut microbiota and TLR4 expression in β-lactoglobulin-induced allergic mice model [2019]
- Additive effect of Lactobacillus acidophilus L-92 on children with atopic dermatitis concomitant with food allergy [2019]
- Efficacy of prolonged ingestion of Lactobacillus acidophilus L-92 in adult patients with atopic dermatitis [2016]
- Effects of oral administration of Lactobacillus acidophilus L-92 on the symptoms and serum cytokines of atopic dermatitis in Japanese adults: a double-blind, randomized, clinical trial [2015]
- [Clinical effects of Lactobacillus acidophilus strain L-55-contained yogurt on symptoms of Japanese cedar pollen allergy] [2012]
- Lactobacillus acidophilus strain L-92 induces CD4(+)CD25(+)Foxp3(+) regulatory T cells and suppresses allergic contact dermatitis [2012]
There are additional studies of these two lactobacillus species with other probiotics being effective with allergies (unfortunately with multiple probiotics being involved, things are unclear as to which part is helping). Note that most lactobacillus species do NOT produce this enzyme.
This is a Novel Approach
The body’s circulating metabolites/chemicals imbalances causing symptoms is well accepted. At present, there is testings of a few of the chemicals that can identify to a medical person that there is an imbalance. They do not provide a clear answer on what the source is. With the use of the microbiome down to the species (and strain) levels, we can estimate levels of over 1600 enzymes. If we have a sufficient reference population, we can identify major shifts. We also jump to an understanding of the root cause — which has eluded us before — our microbiome is not producing enough of certain chemicals due to the current composition of the bacteria population.
In the case of MCAS, it is too little of one specific enzyme. This enzyme happens to be produced by two probiotic species. We have also confirmed that these same two species are known to help allergies (but the exact method of helping is often speculated on).
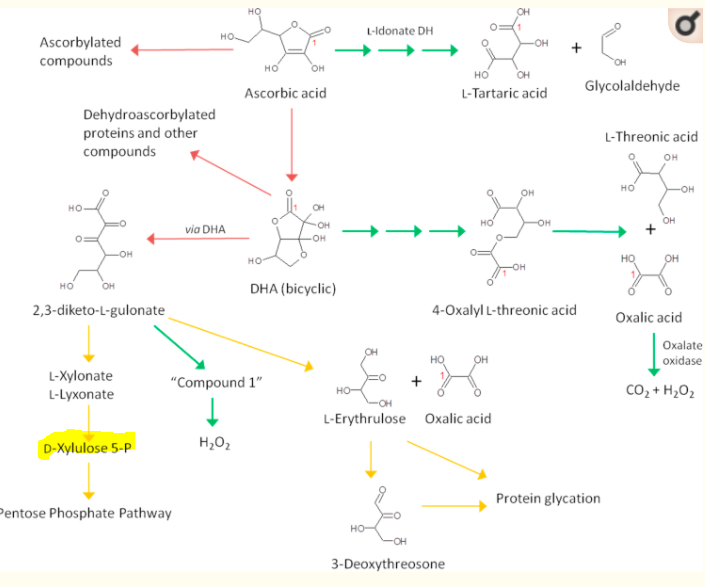
People often want a simple explanation of how it is connected. Unfortunately when dealing with KEGG, we enter the world of a ton of biological wiring as shown by the chart below. This does not means that we can’t use this information; it just means that for most people, the how may be above their formal education and bandwidth.

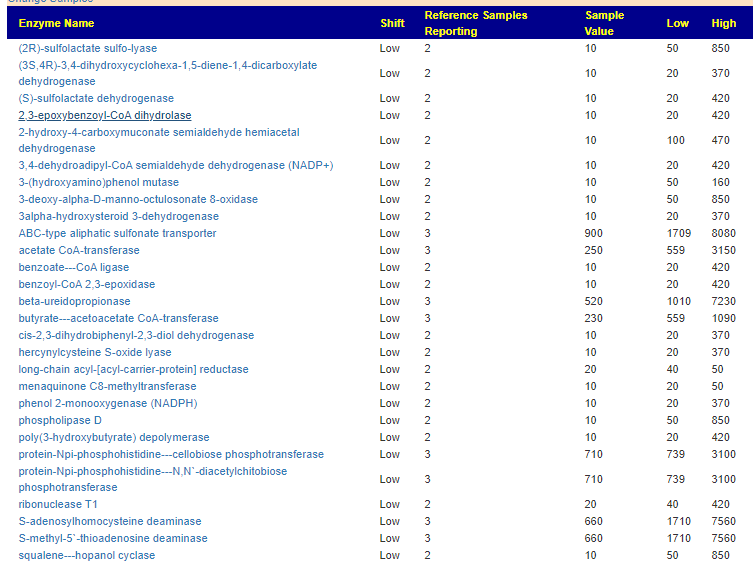
Let us return to this LOW enzyme for a moment and see what is in the literature about it or what is influenced by it: lipopolysaccharide 3-alpha-galactosyltransferase.
- Lipopolysaccharide-mediated mast cell activation induces IFN-gamma secretion by NK cells [2010]
- Lipopolysaccharide suppresses IgE-mast cell-mediated reactions [2017]
- The Mast Cell Is an Early Activator of Lipopolysaccharide-Induced Neuroinflammation and Blood-Brain Barrier Dysfunction in the Hippocampus [2020]
In other words. this enzyme seems connected to mast cells and its activation. Artificial Intelligence magic applied to a large data store of facts, gave us this enlightenment as to cause and suggestions on how to improve MCAS. We lack any case studies — but given the low risk, it is likely worth trying after consulting with your medical professional.
This same pattern can be applied to dozens of individual symptoms (see this page), or across a microbiome sample as a whole.
Addendum
A comment pointed me to Human diamine oxidase is readily released from activated neutrophils ex vivo and in vivo but is rarely elevated in bacteremic patients [Sep 2020] which contains interesting relevant information:
. DAO antigen levels were also determined in three different subcohorts of patients with culture-proven bacteremia and high C-reactive protein (CRP) levels. DAO concentrations were elevated in a time-dependent manner in serum but not in EDTA or citrate plasma (P < 0.01). Neutrophil activation using phorbol myristate acetate (PMA) and zymosan dose-dependently caused DAO concentrations to be elevated more than 10-fold at both 22°C and 37°C (both P-values <0.001). Administration of LPS to healthy volunteers released DAO from neutrophils (P < 0.001). Of the 55 different bacteremic patients selected from three independent cohorts only 3 (5.4%) showed highly elevated DAO concentrations. Serum DAO concentrations do not accurately reflect circulating enzyme levels but coagulation-induced neutrophil activation and consequently DAO release. Only a few bacteremic patients show high DAO concentrations able to degrade histamine rapidly.
Human diamine oxidase is readily released from activated neutrophils ex vivo and in vivo but is rarely elevated in bacteremic patients [Sep 2020]
Lipopolysaccharide are endotoxins. ” Gram negative pathogens may secrete outer membrane vesicles containing lipopolysaccharide endotoxin and some virulence proteins in the bounding membrane along with some other toxins as intra-vesicular contents,” This drops a scent in this direction:
- ” Most microorganisms recognized to date as probiotics are Gram-positive, with Lactobacillus and Bifidobacterium being the main species used as treatments of intestinal dysfunctions (Marco et al. 2006). However, some Gram-negatives are also used as probiotics. The best example of this group is Escherichia coli Nissle 1917 (EcN) (Nissle 1959), also known as “Mutaflor,” which has been used in Germany for many years in the treatment of chronic constipation (Mollenbrink and Bruckschen 1994) and colitis (Schutz 1989).” [2013]
- This leads us to K5 Capsule and Lipopolysaccharide Are Important in Resistance to T4 Phage Attack in Probiotic E. coli Strain Nissle 1917 [2019] which suggests that mutaflor (or an engineered version of it) may be a solution to MCAS. Since Mutaflor takes up residency often, it may be a potential cure. Symbioflor-2 (US Source, World Wide Source) , a different E.Coli probiotic, may also have impact.
It also leads to a hypothesis that could be tested easily: MCAS is associated with lower or non-existent levels of gram-negative bacteria in the microbiome.
CAUTION: What is claimed to be in a probiotic may not be
This is a long standing problem with commercial probiotics — there is no effective regulation that what is advertised is actually in the bottle. In this case, we want specific species — ideally just those species.
My family’s usual source is Custom Probiotics – because their products have no filler, single species usually, and single strains. Unfortunately, they only sell one of these species: Lactobacillus Acidophilus with a recommended adult dosage of 160 BCFU/day. A list of commercial probiotics with claimed species is here, unfortunately L.H. is almost always in mixtures except for one UK source, Metabolics, species LMG 26307 (they also sell Lactobacillus Acidophilus, species LMG 8151) so an ideal single source for a MCAS person in the EU who wish to trial these suggestions.
If you an “off the shelf” probiotic mixture, your odds are better of actually getting the right species when the probiotic does not just say “lactobacillus helveticus” but gives a strain, i.e. one of these: ATCC:15009, CCUG:30139, CIP:103146, DSM:20075 , IFO:15019, JCM:1120, BCCM/LMG:13555, BCCM/LMG:6413, NBRC:15019, NRRL:B-4526, CGMCC:1.1877. It is available from manufacturers, i.e. creative enzymes.
- “A new study by scientists at the University of California has found that contents of many bifidobacteria probiotic products differ from the ingredients listed….16 products.. only one matched the ingredient list .. Some products also contained non-label species” [Source 2015]
- “Current trademark law and the lack of stringent regulation of probiotic manufacturing mean that the trademark owner can commercialize any formulation under the same brand, even if significantly different from the original.” The Unregulated Probiotic Market
- My earlier post: Deceptive Probiotic Labels
Genome Sequence of Lactobacillus helveticus, an Organism Distinguished by Selective Gene Loss and Insertion Sequence Element Expansion tells how unique this species is.