Over this weekend, we finally reached the threshold for citizen science to look at Autism (20+ symptom annotated uploads). This coming weekend I am attending a conference on Autism in Vancouver, British Columbia, Canada where Jason Hawrelak from Tasmania will be speaking (schedule below).
In preparation for this I decided to do a technical study comparing published studies on Autism and the results of citizen science. Is there agreement? What are the factors that come up with different results?

Video Version
Source Data
Data comes from two sources (both are subject to change with time):
- Publish Studies – listed here with citations
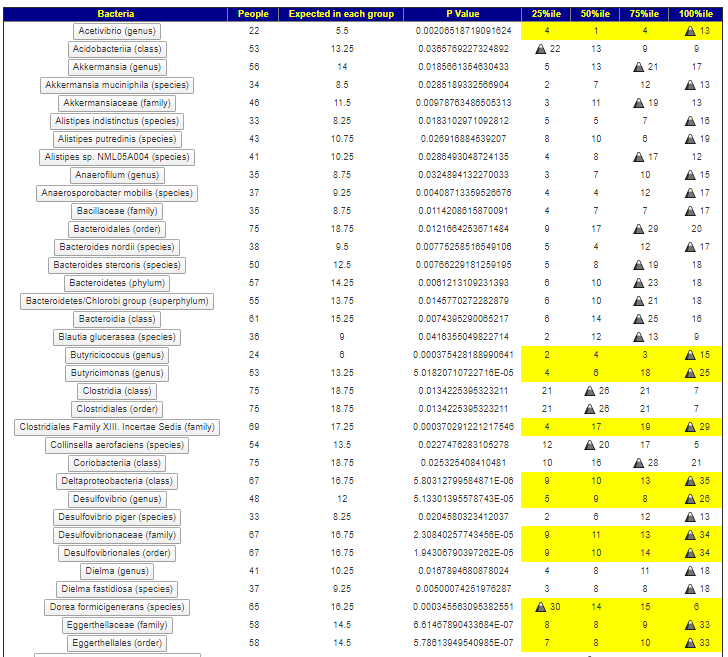
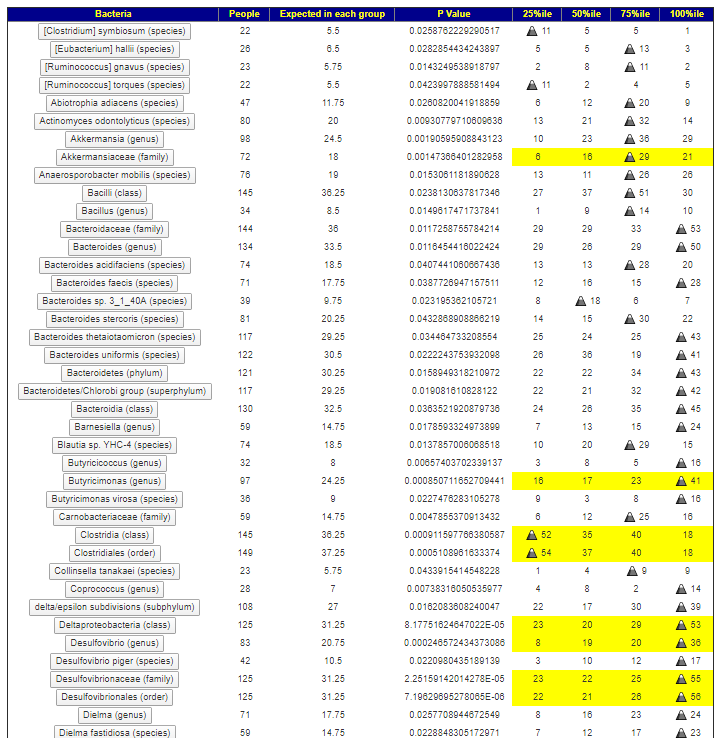
- Citizen Science from Microbiome Prescription – showing the distributions
We had agreement on several items, namely:
- Sutterellaceae (family) being High
- Sutterella (genus) being High
- Blautia being Low
- Bifidobacterium being High (in some Published studies)
- Lactobacillus being High (in some Published studies)
We had disagreement on only one item:
- Veillonella was high on citizen science and low on published studies
Age is a significant factor for both Bifidobacterium and Lactobacillus. Studies on autism under 6, 6-12 and older will have significantly different amounts. This may account for disagreement between published studies.
Critique of Published Studies
Reproducibility of studies is important. There are 17 Pub Med studies on the autism microbiome. If you look at Publish Studies, the same bacteria is only reported 7 times at most. Many bacteria are only reported once. Whether a bacteria is evaluated depends on the equipment being used, for example – Bilophila is reported as significant in 2 studies but Thryve 16s results do not report it.
The usual criteria is to detect statistical significance of averages between the target group and the control group. This requires some major, likely false, assumptions:
- Representative samples in each group
- Normal distribution of the bacteria in each group
A single outlier in a group can cause statistical significance to appear while a more sensitive (but require more effort) test like Pearson’s chi-squared test would find nothing to publish about. It is important to note that Pearson’s usually results in sample sizes of at least 20; many published studies do not have that number.
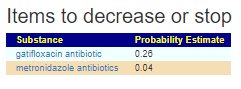
If I exclude non-matches with at least 2 studies with consistent results (a single study in 17 studies is likely unreliable), then we have the following not detected yet with Citizen Science (which have barely enough studies to do any analysis).
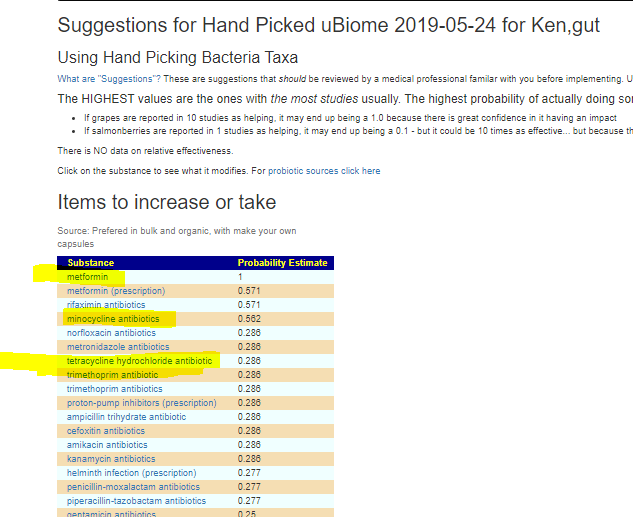
On the flip side, Citizen Science reports on some bacteria not reported in published studies over 10 with high significance.
DNA SNPs and Microbiome Interaction
The microbiome and the person’s DNA interacts. Specific SNP issues are expected to manifest themselves in a person’s microbiome. This adds another level of complexity.

Bottom Line
Large representative samples and in depth skilled statistical analysis is needed to reliably identify the bacteria associated with a symptom, medical condition or characteristic.
From Publish Studies and Citizen Science, we have apparent agreement on 5 taxa, disagreement on 1, and silence in terms of repeatability on some 20 additional ones. I hope/ believe that as the number of samples in Citizen Science data set grows we will find more agreement with some of the published studies.
Below is a spreadsheet for interested parties.
References
- The Gut Microbiota and Autism Spectrum Disorders
Frontiers in Cellular Neuroscience (Front Cell Neurosci ) Vol: 11 Issue Pages: 120
Pub: 2017 Apr 28 Epub: 2017 Apr 28 Authors Li Q , Han Y , Dy AB , Hagerman RJ , - New evidences on the altered gut microbiota in autism spectrum disorders.
Microbiome (Microbiome ) Vol: 5 Issue 1 Pages: 24
Pub: 2017 Feb 22 Epub: 2017 Feb 22 Authors Strati F , Cavalieri D , Albanese D , De Felice C , Donati C , Hayek J , Jousson O , Leoncini S , Renzi D , Calabrò A , De Filippo C , - Distinct Microbiome-Neuroimmune Signatures Correlate With Functional Abdominal Pain in Children With Autism Spectrum Disorder.
Cellular and molecular gastroenterology and hepatology (Cell Mol Gastroenterol Hepatol ) Vol: 3 Issue 2 Pages: 218-230
Pub: 2017 Mar Epub: 2016 Dec 11 Authors Luna RA , Oezguen N , Balderas M , Venkatachalam A , Runge JK , Versalovic J , Veenstra-VanderWeele J , Anderson GM , Savidge T , Williams KC , - The valproic acid rat model of autism presents with gut bacterial dysbiosis similar to that in human autism.
Molecular autism (Mol Autism ) Vol: 9 Issue Pages: 61
Pub: 2018 Epub: 2018 Dec 10 Authors Liu F , Horton-Sparks K , Hull V , Li RW , Martínez-Cerdeño V , - Analysis of gut microbiota profiles and microbe-disease associations in children with autism spectrum disorders in China.
Scientific reports (Sci Rep ) Vol: 8 Issue 1 Pages: 13981
Pub: 2018 Sep 18 Epub: 2018 Sep 18 Authors Zhang M , Ma W , Zhang J , He Y , Wang J , - Fecal microbiota and metabolome of children with autism and pervasive developmental disorder not otherwise specified.
PloS one (PLoS One ) Vol: 8 Issue 10 Pages: e76993
Pub: 2013 Epub: 2013 Oct 9 Authors De Angelis M , Piccolo M , Vannini L , Siragusa S , De Giacomo A , Serrazzanetti DI , Cristofori F , Guerzoni ME , Gobbetti M , Francavilla R , - Identifying psychiatric disorder-associated gut microbiota using microbiota-related gene set enrichment analysis.
Briefings in bioinformatics (Brief Bioinform ) Vol: Issue Pages:
Pub: 2019 Apr 5 Epub: 2019 Apr 5 Authors Cheng S , Han B , Ding M , Wen Y , Ma M , Zhang L , Qi X , Cheng B , Li P , Kafle OP , Liang X , Liu L , Du Y , Zhao Y , Zhang F , - Altered composition and function of intestinal microbiota in autism spectrum disorders: a systematic review.
Translational psychiatry (Transl Psychiatry ) Vol: 9 Issue 1 Pages: 43
Pub: 2019 Jan 29 Epub: 2019 Jan 29 Authors Liu F , Li J , Wu F , Zheng H , Peng Q , Zhou H , - Microbiota-related Changes in Bile Acid & Tryptophan Metabolism are Associated with Gastrointestinal Dysfunction in a Mouse Model of Autism.
EBioMedicine (EBioMedicine ) Vol: 24 Issue Pages: 166-178
Pub: 2017 Oct Epub: 2017 Sep 21 Authors Golubeva AV , Joyce SA , Moloney G , Burokas A , Sherwin E , Arboleya S , Flynn I , Khochanskiy D , Moya-Pérez A , Peterson V , Rea K , Murphy K , Makarova O , Buravkov S , Hyland NP , Stanton C , Clarke G , Gahan CGM , Dinan TG , Cryan JF , - Intestinal Dysbiosis and Yeast Isolation in Stool of Subjects with Autism Spectrum Disorders.
Mycopathologia (Mycopathologia ) Vol: 182 Issue 3-4 Pages: 349-363
Pub: 2017 Apr Epub: 2016 Sep 21 Authors Iovene MR , Bombace F , Maresca R , Sapone A , Iardino P , Picardi A , Marotta R , Schiraldi C , Siniscalco D , Serra N , de Magistris L , Bravaccio C , - Ketogenic diet modifies the gut microbiota in a murine model of autism spectrum disorder.
Molecular autism (Mol Autism ) Vol: 7 Issue 1 Pages: 37
Pub: 2016 Epub: 2016 Sep 1 Authors Newell C , Bomhof MR , Reimer RA , Hittel DS , Rho JM , Shearer J , - Can we reduce autism-related gastrointestinal and behavior problems by gut microbiota based dietary modulation? A review.
Nutritional neuroscience (Nutr Neurosci ) Vol: Issue Pages: 1-12
Pub: 2019 Jun 19 Epub: 2019 Jun 19 Authors Nogay NH , Nahikian-Nelms M , - The Role of Gut Microbiota in Gastrointestinal Symptoms of Children with ASD.
Medicina (Kaunas, Lithuania) (Medicina (Kaunas) ) Vol: 55 Issue 8 Pages:
Pub: 2019 Jul 26 Epub: 2019 Jul 26 Authors Martínez-González AE , Andreo-Martínez P , - Association Between Gut Microbiota and Autism Spectrum Disorder: A Systematic Review and Meta-Analysis.
Frontiers in psychiatry (Front Psychiatry ) Vol: 10 Issue Pages: 473
Pub: 2019 Epub: 2019 Jul 17 Authors Xu M , Xu X , Li J , Li F , - Analysis of gut microbiome, nutrition and immune status in autism spectrum disorder: a case-control study in Ecuador.
Gut microbes (Gut Microbes ) Vol: Issue Pages: 1-12
Pub: 2019 Sep 18 Epub: 2019 Sep 18 Authors Zurita MF , Cárdenas PA , Sandoval ME , Peña MC , Fornasini M , Flores N , Monaco MH , Berding K , Donovan SM , Kuntz T , Gilbert JA , Baldeón ME , - Increased abundance of Sutterella spp. and Ruminococcus torques in feces of children with autism spectrum disorder.
Molecular autism (Mol Autism ) Vol: 4 Issue 1 Pages: 42
Pub: 2013 Nov 4 Epub: 2013 Nov 4 Authors Wang L , Christophersen CT , Sorich MJ , Gerber JP , Angley MT , Conlon MA , - Characterization of Intestinal Microbiota and Probiotics Treatment in Children With Autism Spectrum Disorders in China.
Frontiers in neurology (Front Neurol ) Vol: 10 Issue Pages: 1084
Pub: 2019 Epub: 2019 Nov 5 Authors Niu M , Li Q , Zhang J , Wen F , Dang W , Duan G , Li H , Ruan W , Yang P , Guan C , Tian H , Gao X , Zhang S , Yuan F , Han Y , - Autism spectrum disorder is associated with gut microbiota disorder in children.
BMC pediatrics (BMC Pediatr ) Vol: 19 Issue 1 Pages: 516
Pub: 2019 Dec 27 Epub: 2019 Dec 27 Authors Sun H , You Z , Jia L , Wang F , - An approach to gut microbiota profile in children with autism spectrum disorder.
Environmental microbiology reports (Environ Microbiol Rep ) Vol: Issue Pages:
Pub: 2019 Nov 11 Epub: 2019 Nov 11 Authors Andreo-Martínez P , García-Martínez N , Sánchez-Samper EP , Martínez-González AE , - [Correlation between gut microbiota and behavior symptoms in children with autism spectrum disorder].
Zhongguo dang dai er ke za zhi = Chinese journal of contemporary pediatrics (Zhongguo Dang Dai Er Ke Za Zhi ) Vol: 21 Issue 7 Pages: 663-669
Pub: 2019 Jul Epub: Authors Zhao RH , Zheng PY , Liu SM , Tang YC , Li EY , Sun ZY , Jiang MM ,
For people not aware of the Microbiome Prescription site, this gives an overview.