For any one that is interested, bacteria with P < 0.005 significance to 324 symptoms and diagnosis is now available (with source data) at https://microbiomeprescription.com/sample/Frequency
Some items of interest to the ME/CFS Community are below


For any one that is interested, bacteria with P < 0.005 significance to 324 symptoms and diagnosis is now available (with source data) at https://microbiomeprescription.com/sample/Frequency
Some items of interest to the ME/CFS Community are below



In my last post on MRI Scans, I felt the best model is based on Evidence of widespread metabolite abnormalities in Myalgic encephalomyelitis/chronic fatigue syndrome: assessment with whole-brain magnetic resonance spectroscopy [2020]. Metabolite abnormalities can be a direct result of microbiome dysfunctions. Those abnormalities are very treatable using microbiome tests and expert systems such as generated by Microbiome Prescription.
Metabolites are substances made or used in the body during metabolism, which is the process of breaking down food or chemicals into energy and other useful materials. They help the body grow, repair itself, and function properly. Examples include amino acids, vitamins, and sugars.
In Myalgic Encephalomyelitis/Chronic Fatigue Syndrome (ME/CFS), metabolites have been found to play a critical role in understanding the disease’s mechanisms and symptoms:
These findings highlight the importance of metabolites in ME/CFS research, offering potential biomarkers for diagnosis and targets for therapeutic interventions.
In the context of Irritable Bowel Syndrome (IBS), metabolites play a significant role:
Understanding these metabolites and their interactions with the gut microbiome may provide valuable insights into the underlying mechanisms of IBS and potentially lead to new diagnostic tools or treatments.
Enzymes play a crucial role in managing metabolites within our bodies. Here’s a simple description of their relationship:
In essence, enzymes are the workers that manage metabolites, ensuring our bodies can efficiently use the food we eat and carry out the chemical processes necessary for life.
It happens that from uploaded samples and KEGG: Kyoto Encyclopedia of Genes and Genomes; we can determine that the following enzymes are (VERY VERY) statistically significant. The most significant ones are all too high. The top ones comes from the three genus only: Chlorobaculum , Pelodictyon and Prosthecochloris
Some (but not all) enzymes can be provided by some probiotics. Below is recent feedback from a person dealing with a child’s autism.

| EC Key | Enzyme Name | Probability | Shift |
| 1.1.1.325 | sepiapterin reductase (L–threo-7,8-dihydrobiopterin forming) | 1.61685e-015 | high |
|---|---|---|---|
| 2.1.1.331 | bacteriochlorophyllide d C-121-methyltransferase | 1.61685e-015 | high |
| 2.1.1.332 | bacteriochlorophyllide d C-82-methyltransferase | 1.61685e-015 | high |
| 2.1.1.333 | bacteriochlorophyllide d C-20 methyltransferase | 1.61685e-015 | high |
| 3.1.1.100 | chlorophyllide a hydrolase | 1.61685e-015 | high |
| 4.2.1.169 | 3-vinyl bacteriochlorophyllide d 31-hydratase | 1.61685e-015 | high |
| 2.3.3.8 | ATP citrate synthase | 1.18208e-013 | high |
| 1.3.1.75 | 3,8-divinyl protochlorophyllide a 8-vinyl-reductase (NADPH) | 5.353e-013 | high |
| 2.5.1.42 | geranylgeranylglycerol-phosphate geranylgeranyltransferase | 7.71322e-013 | high |
| 1.17.98.2 | bacteriochlorophyllide c C-71-hydroxylase | 2.69579e-012 | high |
| 2.7.8.36 | undecaprenyl phosphate N,N′-diacetylbacillosamine 1-phosphate transferase | 3.60014e-009 | low |
| 1.11.1.6 | catalase | 1.37633e-008 | low |
| 6.5.1.8 | 3′-phosphate/5′-hydroxy nucleic acid ligase | 1.02548e-007 | low |
| 2.5.1.105 | 7,8-dihydropterin-6-yl-methyl-4-(β-D-ribofuranosyl)aminobenzene 5′-phosphate synthase | 1.27364e-007 | low |
| 1.1.1.65 | pyridoxine 4-dehydrogenase | 1.89183e-007 | high |
| 2.6.1.59 | dTDP-4-amino-4,6-dideoxygalactose transaminase | 3.34948e-007 | low |
| 4.1.1.31 | phosphoenolpyruvate carboxylase | 4.73602e-007 | high |
| 5.4.99.26 | tRNA pseudouridine65 synthase | 5.94186e-007 | high |
| 1.1.1.127 | 2-dehydro-3-deoxy-D-gluconate 5-dehydrogenase | 8.09119e-007 | low |
| 3.4.21.83 | oligopeptidase B | 8.612e-007 | high |
| 1.1.1.9 | D-xylulose reductase | 1.07212e-006 | high |
| 1.12.1.4 | hydrogenase (NAD+, ferredoxin) | 1.22627e-006 | high |
| 3.5.2.9 | 5-oxoprolinase (ATP-hydrolysing) | 1.30156e-006 | high |
| 2.7.1.12 | gluconokinase | 1.49961e-006 | high |
| 1.6.1.2 | NAD(P)+ transhydrogenase (Re/Si-specific) | 3.05641e-006 | high |
| 7.1.1.1 | proton-translocating NAD(P)+ transhydrogenase | 3.05641e-006 | high |
| 3.1.1.114 | methyl acetate hydrolase | 3.98055e-006 | low |
| 3.2.1.165 | exo-1,4-β-D-glucosaminidase | 4.31375e-006 | low |
| 2.3.1.117 | 2,3,4,5-tetrahydropyridine-2,6-dicarboxylate N-succinyltransferase | 4.31566e-006 | high |
| 2.7.8.12 | teichoic acid poly(glycerol phosphate) polymerase | 4.59165e-006 | low |
| 6.1.1.13 | D-alanine—poly(phosphoribitol) ligase | 4.88544e-006 | low |
| 1.12.98.4 | sulfhydrogenase | 6.48299e-006 | low |
| 4.2.1.22 | cystathionine β-synthase | 7.49383e-006 | high |
| 2.3.1.78 | heparan-α-glucosaminide N-acetyltransferase | 8.1071e-006 | low |
This is an update of my post from 10 years ago, CFS: Appropriate Brain Scans. I will focus on studies in those 10 years. Short version of these studies below.
Data showed that MRI studies frequently reported structural changes in the white and gray matter. Abnormalities of the functional connectivity within the brainstem and with other brain regions have also been found. The studies have suggested possible mechanisms including astrocyte dysfunction, cerebral perfusion impairment, impaired nerve conduction, and neuroinflammation involving the brainstem, which may at least partially explain a substantial portion of the ME/CFS symptoms and their heterogeneous presentations in individual patient
Brainstem Abnormalities in Myalgic Encephalomyelitis/Chronic Fatigue Syndrome: A Scoping Review and Evaluation of Magnetic Resonance Imaging Findings [2021]
In our family dealing with ME/CFS, we were fortunate is having brain scans done by and interpreted by Doctor Daniel Amen. Our effective treatment was focused on shifting bacteria, addressing coagulation, and reducing inflammation.
For more information on metabolites, see this post.
I believe the best model is based on Evidence of widespread metabolite abnormalities in Myalgic encephalomyelitis/chronic fatigue syndrome: assessment with whole-brain magnetic resonance spectroscopy [2020]. Metabolite abnormalities can be a direct result of microbiome dysfunctions. Those abnormalities are very treatable using microbiome tests and expert systems such as generated by Microbiome Prescription.
The process is very simple, for a condition like ME/CFS, we compute the expected number of samples reporting this bacteria (based on people without Long COVID) and compare it to the actual number seen. This can be used to compute a statistical value called Chi-Square (χ²), This is then used to compute the chance of it happening at random. This is possible because we have over 3600 samples from some labs and thus able to detect things better.
Actual example:
Biomesight and Ombre identifies bacteria using different methodologies so often give different names and amounts. For background on this lack of standardization, see The taxonomy nightmare before Christmas…
The data below is for samples marked with “Official Diagnosis: Chronic Fatigue Syndrome (CFS/ME)“
For Long COVID, see this post.
Unlike some conditions shown below, it is not just one bacteria involved but combinations.
We have 10 bacteria that are too low and some 67 that are too high. Six of the 10 that are too low should be available as probiotics (conceptually), but only B.Bifidum is.
In the too high group are some familiar names: Rickettsia (Cecile Jadin’s protocol) and spotted fever group.
| Bacteria Name | Rank | Expected | Observed | Shift | Prob |
| Anaerococcus murdochii | species | 22.1 | 40 | Too High | 0.000134 |
| Anaerococcus octavius | species | 23.2 | 36 | Too High | 0.00785 |
| Anaerococcus senegalensis | species | 21.6 | 34 | Too High | 0.007766 |
| Bacteroides fluxus | species | 82.2 | 111 | Too High | 0.001504 |
| Bacteroides thetaiotaomicron | species | 95.6 | 122 | Too High | 0.006947 |
| Bifidobacterium asteroides | species | 50.3 | 28 | Too Low | 0.004406 |
| Bifidobacterium bifidum | species | 76.3 | 49 | Too Low | 0.002295 |
| Bifidobacterium bombi | species | 80.8 | 44 | Too Low | 0.000136 |
| Bifidobacterium commune | species | 56.8 | 30 | Too Low | 0.000581 |
| Bifidobacterium magnum | species | 73.0 | 46 | Too Low | 0.002755 |
| Bifidobacterium thermacidophilum | species | 53.8 | 31 | Too Low | 0.004352 |
| Campylobacter | genus | 54.0 | 74 | Too High | 0.006445 |
| Campylobacter hominis | species | 26.8 | 43 | Too High | 0.001707 |
| Campylobacteraceae | family | 54.8 | 76 | Too High | 0.004245 |
| Campylobacterales | order | 61.6 | 83 | Too High | 0.006328 |
| Desulfarculaceae | family | 24.9 | 40 | Too High | 0.002442 |
| Desulfarculales | order | 24.9 | 40 | Too High | 0.002442 |
| Desulfarculia | class | 26.2 | 40 | Too High | 0.00692 |
| Desulfocarbo | genus | 14.8 | 26 | Too High | 0.003442 |
| Desulfocarbo indianensis | species | 14.6 | 26 | Too High | 0.002722 |
| Desulfoglaeba | genus | 27.8 | 42 | Too High | 0.007259 |
| Desulfoglaeba alkanexedens | species | 27.4 | 42 | Too High | 0.005339 |
| Desulfovibrio legallii | species | 32.5 | 48 | Too High | 0.006443 |
| Desulfovibrio piger | species | 35.9 | 53 | Too High | 0.004287 |
| Dialister pneumosintes | species | 14.8 | 27 | Too High | 0.001445 |
| Emticicia | genus | 19.3 | 33 | Too High | 0.001758 |
| Emticicia sediminis | species | 8.4 | 21 | Too High | 1.52E-05 |
| Enorma | genus | 22.4 | 35 | Too High | 0.007472 |
| Enorma massiliensis | species | 22.4 | 35 | Too High | 0.007472 |
| Enterorhabdus | genus | 69.6 | 45 | Too Low | 0.004671 |
| Epsilonproteobacteria | class | 62.7 | 84 | Too High | 0.007016 |
| Ezakiella | genus | 44.7 | 63 | Too High | 0.006216 |
| Flavobacteriaceae | family | 75.1 | 98 | Too High | 0.008138 |
| Halanaerobiales | order | 78.4 | 54 | Too Low | 0.005779 |
| Hallella | genus | 34.4 | 60 | Too High | 1.30E-05 |
| Hallella bergensis | species | 17.5 | 39 | Too High | 2.77E-07 |
| Hallella multisaccharivorax | species | 26.2 | 45 | Too High | 0.000235 |
| Holdemania massiliensis | species | 55.2 | 75 | Too High | 0.00788 |
| Hoylesella buccalis | species | 75.1 | 105 | Too High | 0.000552 |
| Hoylesella loescheii | species | 34.3 | 51 | Too High | 0.004268 |
| Hoylesella marshii | species | 15.6 | 30 | Too High | 0.000268 |
| Hoylesella nanceiensis | species | 30.4 | 48 | Too High | 0.001467 |
| Humidesulfovibrio | genus | 29.9 | 49 | Too High | 0.000497 |
| Humidesulfovibrio idahonensis | species | 27.8 | 47 | Too High | 0.000281 |
| Hungateiclostridiaceae | family | 78.0 | 106 | Too High | 0.001492 |
| Insolitispirillum | genus | 31.0 | 46 | Too High | 0.007226 |
| Insolitispirillum peregrinum | species | 30.4 | 46 | Too High | 0.004822 |
| Leptolinea | genus | 14.4 | 26 | Too High | 0.002277 |
| Leptolinea tardivitalis | species | 13.5 | 24 | Too High | 0.004247 |
| Parabacteroides johnsonii | species | 51.0 | 70 | Too High | 0.007925 |
| Paraprevotella xylaniphila | species | 25.0 | 42 | Too High | 0.000677 |
| Parvimonas | genus | 29.7 | 52 | Too High | 4.33E-05 |
| Parvimonas micra | species | 29.7 | 52 | Too High | 4.33E-05 |
| Pedobacter | genus | 36.3 | 53 | Too High | 0.005472 |
| Peptoniphilus lacrimalis | species | 34.6 | 51 | Too High | 0.005246 |
| Persicobacteraceae | family | 12.1 | 24 | Too High | 0.000587 |
| Phocaeicola salanitronis | species | 105.9 | 136 | Too High | 0.003396 |
| Porphyromonas asaccharolytica | species | 33.8 | 51 | Too High | 0.003157 |
| Porphyromonas endodontalis | species | 25.7 | 40 | Too High | 0.004944 |
| Prevotella dentasini | species | 6.8 | 29 | Too High | 1.25E-17 |
| Prevotella denticola | species | 4.2 | 27 | Too High | 1.35E-28 |
| Prolixibacteraceae | family | 20.7 | 33 | Too High | 0.006663 |
| Propioniferax | genus | 23.2 | 42 | Too High | 9.45E-05 |
| Propioniferax innocua | species | 22.8 | 42 | Too High | 5.61E-05 |
| Rickettsia | genus | 21.0 | 36 | Too High | 0.001101 |
| Rickettsia honei | species | 14.8 | 28 | Too High | 0.000569 |
| Segatella bryantii | species | 24.5 | 39 | Too High | 0.003287 |
| Segatella maculosa | species | 33.2 | 55 | Too High | 0.000161 |
| Segatella oris | species | 22.2 | 38 | Too High | 0.000807 |
| Segatella paludivivens | species | 32.9 | 54 | Too High | 0.000245 |
| Senegalimassilia anaerobia | species | 54.0 | 34 | Too Low | 0.006868 |
| Slackia piriformis | species | 56.5 | 29 | Too Low | 0.00048 |
| spotted fever group | species group | 17.9 | 32 | Too High | 0.000907 |
| Tindallia | genus | 14.8 | 26 | Too High | 0.003442 |
| unclassified Burkholderiales | family | 20.9 | 33 | Too High | 0.008034 |
| unclassified Clostridiales | family | 77.1 | 106 | Too High | 0.000984 |
We have more data from Biomesight which means better (more) detection of significant bacteria.
As above, we have 12 bacteria that are too low and 116 bacteria that are too high. We have only 2 Bifidobacterium identified with only one available as a probiotic
We see that Lactobacillus is too high. This agrees with brain fog being caused by over production of d-lactic acid.
We also see these two are high that are commonly associated to issues:
| Tax_Name | Tax_Rank | Expected | Observed | Shift | Probability |
| [Actinobacillus] rossii | species | 119.1 | 89 | Too Low | 0.005858 |
| [Pasteurella] aerogenes-[Pasteurella] mairii-[Actinobacillus] rossii complex | species group | 119.1 | 89 | Too Low | 0.005858 |
| Absiella | genus | 29.8 | 56 | Too High | 1.52E-06 |
| Acetobacteraceae | family | 103.7 | 131 | Too High | 0.007414 |
| Acholeplasma hippikon | species | 73.9 | 110 | Too High | 2.63E-05 |
| Actinobacillus | genus | 176.6 | 134 | Too Low | 0.004701 |
| Actinobacillus pleuropneumoniae | species | 61.6 | 35 | Too Low | 0.00299 |
| Actinobacillus porcinus | species | 133.9 | 99 | Too Low | 0.003018 |
| Adlercreutzia equolifaciens | species | 189.7 | 227 | Too High | 0.006755 |
| Aggregatibacter | genus | 82.3 | 49 | Too Low | 0.000912 |
| Alcanivoracaceae | family | 37.3 | 54 | Too High | 0.006427 |
| Alcanivorax | genus | 37.3 | 54 | Too High | 0.006427 |
| Amedibacillus | genus | 196.8 | 235 | Too High | 0.006495 |
| Amedibacillus dolichus | species | 197.1 | 235 | Too High | 0.006912 |
| Anaerococcus | genus | 126.4 | 157 | Too High | 0.006444 |
| Anaerococcus hydrogenalis | species | 39.1 | 58 | Too High | 0.002475 |
| Anaerococcus prevotii | species | 21.1 | 36 | Too High | 0.001188 |
| Anaerococcus tetradius | species | 29.6 | 59 | Too High | 6.37E-08 |
| Anaerococcus vaginalis | species | 63.6 | 87 | Too High | 0.003269 |
| Anaerofustis | genus | 54.3 | 76 | Too High | 0.003316 |
| Anaerofustis stercorihominis | species | 53.0 | 74 | Too High | 0.003981 |
| Arthrobacter | genus | 27.7 | 43 | Too High | 0.003655 |
| Bifidobacterium adolescentis | strain | 102.4 | 63 | Too Low | 0.001744 |
| Bifidobacterium cuniculi | species | 64.9 | 38 | Too Low | 0.007829 |
| Bifidobacterium ruminantium | species | 14.8 | 30 | Too High | 7.46E-05 |
| Caloramator viterbiensis | species | 19.0 | 37 | Too High | 3.60E-05 |
| Campylobacter ureolyticus | species | 38.0 | 56 | Too High | 0.003481 |
| Cetobacterium | genus | 17.7 | 29 | Too High | 0.007331 |
| Cetobacterium ceti | species | 13.7 | 28 | Too High | 0.000115 |
| Chlorobaculum limnaeum | species | 8.6 | 22 | Too High | 4.66E-06 |
| Clostridium cadaveris | species | 106.5 | 140 | Too High | 0.001155 |
| Collinsella tanakaei | species | 43.8 | 62 | Too High | 0.005945 |
| Corynebacteriaceae | family | 133.0 | 164 | Too High | 0.007125 |
| Corynebacterium | genus | 133.0 | 164 | Too High | 0.007125 |
| Corynebacterium amycolatum | species | 20.6 | 36 | Too High | 0.000675 |
| Corynebacterium glucuronolyticum | species | 10.4 | 21 | Too High | 0.001029 |
| Corynebacterium pyruviciproducens | species | 12.7 | 23 | Too High | 0.003812 |
| Corynebacterium xerosis | species | 14.8 | 25 | Too High | 0.007807 |
| Dehalobacterium | genus | 124.4 | 159 | Too High | 0.001898 |
| Dehalococcoidaceae | family | 11.9 | 23 | Too High | 0.00124 |
| Dehalococcoidales | order | 11.9 | 23 | Too High | 0.00124 |
| Dehalococcoidia | class | 11.9 | 23 | Too High | 0.00124 |
| Dehalogenimonas | genus | 11.9 | 23 | Too High | 0.00124 |
| Dehalogenimonas lykanthroporepellens | species | 11.3 | 23 | Too High | 0.000539 |
| Desulfonatronovibrio | genus | 44.2 | 62 | Too High | 0.007398 |
| Desulfonatronovibrionaceae | family | 43.3 | 62 | Too High | 0.004437 |
| Desulfovibrio psychrotolerans | species | 19.8 | 37 | Too High | 0.000109 |
| Dethiobacteraceae | family | 47.8 | 77 | Too High | 2.50E-05 |
| Dethiobacterales | order | 47.8 | 77 | Too High | 2.50E-05 |
| Dethiobacteria | class | 47.8 | 77 | Too High | 2.50E-05 |
| Dyadobacter | genus | 77.6 | 103 | Too High | 0.003878 |
| Eggerthella sinensis | species | 133.2 | 167 | Too High | 0.003441 |
| Erysipelothrix inopinata | species | 42.5 | 62 | Too High | 0.002739 |
| Ethanoligenens | genus | 158.8 | 205 | Too High | 0.000248 |
| Eubacterium limosum | species | 27.2 | 42 | Too High | 0.00458 |
| Facklamia | genus | 21.2 | 34 | Too High | 0.005361 |
| Filifactor alocis | species | 35.1 | 54 | Too High | 0.001384 |
| Finegoldia magna | species | 112.9 | 143 | Too High | 0.004643 |
| Halanaerobiaceae | family | 56.2 | 78 | Too High | 0.003557 |
| Halanaerobiales | order | 61.2 | 83 | Too High | 0.005272 |
| Halanaerobium | genus | 56.2 | 78 | Too High | 0.003557 |
| Haploplasma | genus | 13.9 | 31 | Too High | 4.32E-06 |
| Haploplasma cavigenitalium | species | 13.9 | 31 | Too High | 4.32E-06 |
| Heliorestis baculata | species | 17.4 | 30 | Too High | 0.002634 |
| Hungateiclostridiaceae | family | 45.4 | 71 | Too High | 0.000143 |
| Hungateiclostridium | genus | 45.2 | 71 | Too High | 0.000124 |
| Hyphomicrobium | genus | 39.8 | 58 | Too High | 0.004009 |
| Hyphomicrobium aestuarii | species | 34.0 | 51 | Too High | 0.003635 |
| Hyphomonadaceae | family | 16.9 | 29 | Too High | 0.003196 |
| Hyphomonadales | order | 16.9 | 29 | Too High | 0.003196 |
| Isoalcanivorax | genus | 35.5 | 52 | Too High | 0.005689 |
| Isoalcanivorax indicus | species | 31.1 | 52 | Too High | 0.000184 |
| Lactobacillus acidophilus | species | 18.8 | 32 | Too High | 0.002356 |
| Lactobacillus iners | species | 17.3 | 32 | Too High | 0.000436 |
| Legionella shakespearei | species | 67.0 | 91 | Too High | 0.003402 |
| Levilactobacillus | genus | 12.2 | 25 | Too High | 0.000263 |
| Listeriaceae | family | 178.9 | 215 | Too High | 0.006915 |
| Luteolibacter | genus | 127.8 | 159 | Too High | 0.005844 |
| Luteolibacter algae | species | 126.9 | 158 | Too High | 0.005805 |
| Mannheimia | genus | 134.7 | 102 | Too Low | 0.004859 |
| Mannheimia caviae | species | 128.7 | 97 | Too Low | 0.005142 |
| Meiothermus | genus | 78.6 | 103 | Too High | 0.005964 |
| Meiothermus granaticius | species | 77.3 | 101 | Too High | 0.007138 |
| Mobiluncus | genus | 31.7 | 52 | Too High | 0.0003 |
| Mogibacterium vescum | species | 57.6 | 94 | Too High | 1.64E-06 |
| Nannocystineae | suborder | 23.0 | 36 | Too High | 0.006465 |
| Olivibacter terrae | species | 12.1 | 22 | Too High | 0.004168 |
| Paenibacillus filicis | species | 22.9 | 38 | Too High | 0.001632 |
| Pasteurellaceae incertae sedis | no rank | 119.1 | 89 | Too Low | 0.005858 |
| Pediococcus | genus | 166.2 | 208 | Too High | 0.00119 |
| Pediococcus argentinicus | species | 17.9 | 31 | Too High | 0.002047 |
| Peptoniphilus asaccharolyticus | species | 116.8 | 148 | Too High | 0.003873 |
| Peptostreptococcus stomatis | species | 52.0 | 73 | Too High | 0.003541 |
| Porphyromonas asaccharolytica | species | 70.2 | 101 | Too High | 0.000234 |
| Prosthecobacter | genus | 32.7 | 54 | Too High | 0.000194 |
| Prosthecobacter fluviatilis | species | 18.1 | 33 | Too High | 0.00045 |
| Rhizobiaceae | family | 80.9 | 105 | Too High | 0.007378 |
| Roseomonas | genus | 22.3 | 38 | Too High | 0.000867 |
| Roseospira | genus | 54.9 | 78 | Too High | 0.0018 |
| Ruminiclostridium cellulolyticum | species | 107.1 | 135 | Too High | 0.007053 |
| Segatella paludivivens | species | 72.9 | 45 | Too Low | 0.001097 |
| Slackia faecicanis | species | 98.6 | 128 | Too High | 0.003086 |
| Sporosarcina | genus | 47.9 | 69 | Too High | 0.00235 |
| Sporosarcina pasteurii | species | 46.9 | 69 | Too High | 0.001278 |
| Streptococcus agalactiae | species | 11.9 | 23 | Too High | 0.00124 |
| Streptococcus mutans | species | 31.7 | 51 | Too High | 0.0006 |
| Thauera terpenica | species | 20.6 | 33 | Too High | 0.006179 |
| Thermaceae | family | 94.7 | 125 | Too High | 0.001859 |
| Thermales | order | 94.7 | 125 | Too High | 0.001859 |
| Thermoanaerobacter | genus | 78.4 | 102 | Too High | 0.007565 |
| Thermodesulfatator | genus | 22.7 | 42 | Too High | 5.03E-05 |
| Thermodesulfatator atlanticus | species | 22.7 | 42 | Too High | 5.03E-05 |
| Thermodesulfatatoraceae | family | 27.5 | 42 | Too High | 0.005627 |
| Thermodesulfobacteria | class | 22.7 | 42 | Too High | 5.03E-05 |
| Thermodesulfobacteriaceae | family | 22.7 | 42 | Too High | 5.03E-05 |
| Thermodesulfobacteriales | order | 22.7 | 42 | Too High | 5.03E-05 |
| Thermosediminibacteraceae | family | 14.2 | 26 | Too High | 0.001847 |
| Thermovenabulum | genus | 14.2 | 26 | Too High | 0.001847 |
| Thermovenabulum ferriorganovorum | species | 14.2 | 26 | Too High | 0.001847 |
| Thermus | genus | 28.5 | 43 | Too High | 0.006576 |
| Thiocapsa | genus | 75.7 | 104 | Too High | 0.001154 |
| unclassified Burkholderiales | family | 15.2 | 29 | Too High | 0.000429 |
| Vagococcus teuberi | species | 47.0 | 66 | Too High | 0.005467 |
| Varibaculum | genus | 47.5 | 73 | Too High | 0.000214 |
| Varibaculum cambriense | species | 19.3 | 32 | Too High | 0.003695 |
| Winkia | genus | 11.0 | 23 | Too High | 0.000324 |
| Winkia neuii | species | 11.0 | 23 | Too High | 0.000324 |
While the company no longer exists, we have a significant number of samples from them.
As above, we have Lactobacillus crispatus being too high. We have just 2 bacteria being too low and 96 being too high.
| Tax_Name | Tax_Rank | Expected | Observed | Shift | Probability |
| Acetitomaculum | genus | 24.4 | 44 | Too High | 7.07E-05 |
| Acholeplasmatales | order | 10.0 | 21 | Too High | 0.000503 |
| Actinomyces sp. 2002-2301122 | species | 14.4 | 25 | Too High | 0.00497 |
| Aggregatibacter | genus | 42.2 | 24 | Too Low | 0.005171 |
| Alistipes indistinctus | species | 62.8 | 90 | Too High | 0.000602 |
| Alistipes sp. NML05A004 | species | 81.8 | 111 | Too High | 0.001227 |
| Alistipes sp. RMA 9912 | species | 13.7 | 25 | Too High | 0.002304 |
| Anaerococcus murdochii | species | 30.5 | 56 | Too High | 3.93E-06 |
| Anaeroplasmataceae | family | 22.8 | 38 | Too High | 0.00148 |
| Anaeroplasmatales | order | 23.1 | 39 | Too High | 0.000914 |
| Archaea | superkingdom | 39.7 | 73 | Too High | 1.32E-07 |
| Atopobium | genus | 17.8 | 30 | Too High | 0.00374 |
| Bacteroides eggerthii | species | 30.5 | 52 | Too High | 9.98E-05 |
| Bacteroides sp. 35AE37 | species | 66.0 | 92 | Too High | 0.00139 |
| Bacteroides sp. XB12B | species | 39.7 | 66 | Too High | 3.10E-05 |
| Blautia stercoris | species | 37.2 | 55 | Too High | 0.003458 |
| Butyricimonas | genus | 88.4 | 114 | Too High | 0.006407 |
| Butyricimonas faecihominis | species | 57.7 | 79 | Too High | 0.005005 |
| Butyricimonas sp. 214-4 | species | 53.8 | 75 | Too High | 0.003921 |
| Butyricimonas virosa | species | 42.3 | 62 | Too High | 0.002455 |
| Caldicoprobacter | genus | 23.9 | 39 | Too High | 0.001957 |
| Caldicoprobacteraceae | family | 44.2 | 68 | Too High | 0.00034 |
| Campylobacter hominis | species | 27.9 | 42 | Too High | 0.007781 |
| Campylobacter ureolyticus | species | 24.4 | 41 | Too High | 0.000762 |
| Catenibacterium | genus | 57.9 | 80 | Too High | 0.003678 |
| Christensenella minuta | species | 23.1 | 36 | Too High | 0.007121 |
| Cloacibacillus | genus | 19.8 | 38 | Too High | 4.36E-05 |
| Cloacibacillus evryensis | species | 14.7 | 31 | Too High | 2.24E-05 |
| Coprobacter secundus | species | 32.8 | 53 | Too High | 0.000426 |
| Corynebacterium diphtheriae | species | 11.5 | 21 | Too High | 0.005334 |
| Corynebacterium sp. 713182/2012 | species | 16.2 | 27 | Too High | 0.006945 |
| Desulfovibrio | genus | 72.6 | 97 | Too High | 0.004103 |
| Desulfovibrio sp. 3_1_syn3 | species | 9.0 | 30 | Too High | 2.23E-12 |
| Dialister micraerophilus | species | 9.6 | 24 | Too High | 3.85E-06 |
| Dialister propionicifaciens | species | 41.0 | 58 | Too High | 0.008017 |
| Dialister sp. S7MSR5 | species | 16.8 | 29 | Too High | 0.002792 |
| Eubacterium | genus | 27.2 | 46 | Too High | 0.000305 |
| Euryarchaeota | phylum | 39.7 | 73 | Too High | 1.32E-07 |
| Fibrobacter | genus | 14.9 | 27 | Too High | 0.001656 |
| Fibrobacteraceae | family | 24.1 | 47 | Too High | 3.08E-06 |
| Fibrobacterales | order | 32.6 | 61 | Too High | 6.22E-07 |
| Fibrobacteria | class | 32.6 | 61 | Too High | 6.22E-07 |
| Fibrobacterota | phylum | 32.6 | 61 | Too High | 6.22E-07 |
| Gelria | genus | 37.6 | 65 | Too High | 7.75E-06 |
| Granulicatella adiacens | species | 69.5 | 45 | Too Low | 0.006077 |
| Herbaspirillum | genus | 46.7 | 72 | Too High | 0.000218 |
| Herbaspirillum seropedicae | species | 39.0 | 62 | Too High | 0.000225 |
| Hespellia | genus | 96.4 | 123 | Too High | 0.006732 |
| Hoylesella timonensis | species | 23.4 | 40 | Too High | 0.000577 |
| Hungateiclostridiaceae | family | 26.4 | 43 | Too High | 0.001246 |
| Hydrogenoanaerobacterium | genus | 40.6 | 64 | Too High | 0.000246 |
| Lactobacillus crispatus | species | 26.1 | 40 | Too High | 0.006759 |
| Lentisphaeria | class | 53.6 | 75 | Too High | 0.003432 |
| Lentisphaerota | phylum | 53.6 | 75 | Too High | 0.003432 |
| Methanobacteria | class | 37.4 | 73 | Too High | 6.10E-09 |
| Methanobacteriaceae | family | 37.4 | 73 | Too High | 6.10E-09 |
| Methanobacteriales | order | 37.4 | 73 | Too High | 6.10E-09 |
| Methanobrevibacter | genus | 36.9 | 71 | Too High | 2.03E-08 |
| Methanobrevibacter smithii | species | 36.4 | 66 | Too High | 9.34E-07 |
| Methanomada group | clade | 37.4 | 73 | Too High | 6.10E-09 |
| Mollicutes | class | 30.0 | 49 | Too High | 0.00052 |
| Murdochiella | genus | 42.2 | 65 | Too High | 0.000434 |
| Mycoplasmatota | phylum | 30.0 | 49 | Too High | 0.00052 |
| Opitutia | class | 16.4 | 33 | Too High | 4.20E-05 |
| Oxalobacteraceae | family | 47.7 | 73 | Too High | 0.000257 |
| Parabacteroides johnsonii | species | 18.8 | 34 | Too High | 0.000451 |
| Parasporobacterium paucivorans | species | 16.2 | 29 | Too High | 0.001388 |
| Parvibacter | genus | 14.1 | 33 | Too High | 4.82E-07 |
| Parvibacter caecicola | species | 12.3 | 29 | Too High | 1.95E-06 |
| Peptococcus | genus | 71.8 | 100 | Too High | 0.000867 |
| Peptoniphilus lacrimalis | species | 30.3 | 46 | Too High | 0.004193 |
| Porphyromonas sp. 2024b | species | 12.7 | 29 | Too High | 4.76E-06 |
| Propionibacteriaceae | family | 18.8 | 31 | Too High | 0.004859 |
| Propionibacteriales | order | 18.8 | 31 | Too High | 0.004859 |
| Puniceicoccales | order | 16.2 | 33 | Too High | 2.76E-05 |
| Robinsoniella | genus | 30.3 | 50 | Too High | 0.00033 |
| Robinsoniella sp. KNHs210 | species | 22.6 | 37 | Too High | 0.002366 |
| Staphylococcaceae | family | 38.5 | 57 | Too High | 0.002786 |
| Staphylococcus | genus | 38.2 | 57 | Too High | 0.00235 |
| Synergistaceae | family | 41.8 | 72 | Too High | 2.96E-06 |
| Synergistales | order | 41.8 | 72 | Too High | 2.96E-06 |
| Synergistia | class | 41.8 | 72 | Too High | 2.96E-06 |
| Synergistota | phylum | 41.8 | 72 | Too High | 2.96E-06 |
| Thermoanaerobacteraceae | family | 37.7 | 65 | Too High | 8.62E-06 |
| Thermoanaerobacterales | order | 39.7 | 66 | Too High | 3.10E-05 |
| unclassified Butyricimonas | no rank | 53.1 | 77 | Too High | 0.001019 |
| unclassified Desulfovibrio | no rank | 12.6 | 43 | Too High | 8.86E-18 |
| unclassified Finegoldia | no rank | 31.8 | 47 | Too High | 0.006982 |
| unclassified Phascolarctobacterium | no rank | 11.5 | 23 | Too High | 0.000738 |
| unclassified Porphyromonas | no rank | 24.4 | 38 | Too High | 0.005801 |
| unclassified Prevotella | no rank | 9.6 | 21 | Too High | 0.000258 |
| unclassified Robinsoniella | no rank | 22.6 | 37 | Too High | 0.002366 |
| unclassified Staphylococcus | no rank | 26.4 | 43 | Too High | 0.001246 |
| Varibaculum | genus | 48.5 | 70 | Too High | 0.001966 |
| Varibaculum sp. CCUG 45114 | species | 36.1 | 54 | Too High | 0.002986 |
| Victivallaceae | family | 53.6 | 75 | Too High | 0.003432 |
| Victivallales | order | 53.6 | 75 | Too High | 0.003432 |
| Victivallis | genus | 53.1 | 75 | Too High | 0.002607 |
In terms of probiotics, there are two that should be considered:
And all Lactobacillus probiotics avoided.
The above information will be eventually integrated into Microbiome Prescription suggestions expert system. The purpose is to first identify the bacteria of concern.
The following bacteria were reported by 2 or 3 of the above
| Aggregatibacter | genus |
| Anaerococcus murdochii | species |
| Campylobacter hominis | species |
| Campylobacter ureolyticus | species |
| Halanaerobiales | order |
| Hungateiclostridiaceae | family |
| Parabacteroides johnsonii | species |
| Peptoniphilus lacrimalis | species |
| Porphyromonas asaccharolytica | species |
| Segatella paludivivens | species |
| unclassified Burkholderiales | family |
| Varibaculum | genus |
Recently I have seen advertisements for smart watches for ME/CFS. The price would be challenging for many people. I suspect that it also has a poor return for the costs.
This post is about the watch family that my wife and I have been using for a few years. The current version is below. On occasion, I have seen the price under $20.00

People who know me, knows that I prefer to work off objective data – for example microbiome test results, or actual physical measurements. This watch has empowered me. It can do automatic measurements every 10 minutes for day long monitoring.
You get an overview of the many measurements on a dashboard on your smart phone.
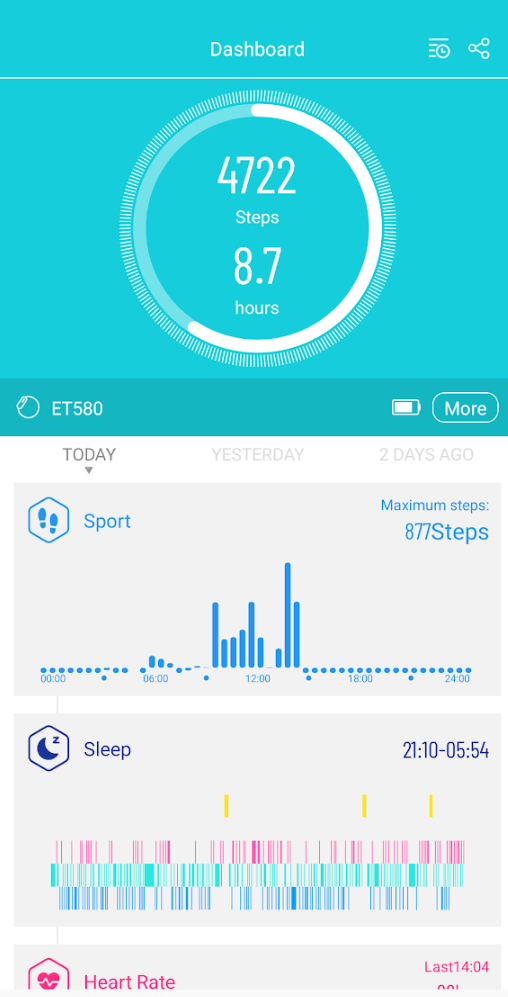

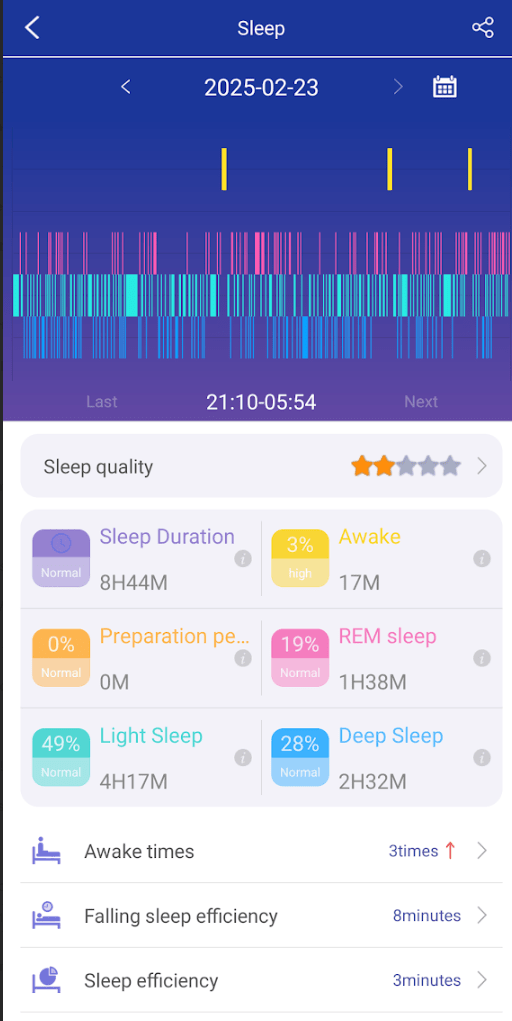
Sleep is often a challenge. My wife found that some supplements improves her sleep and others do not. This changes decision making from being subjective to objective.
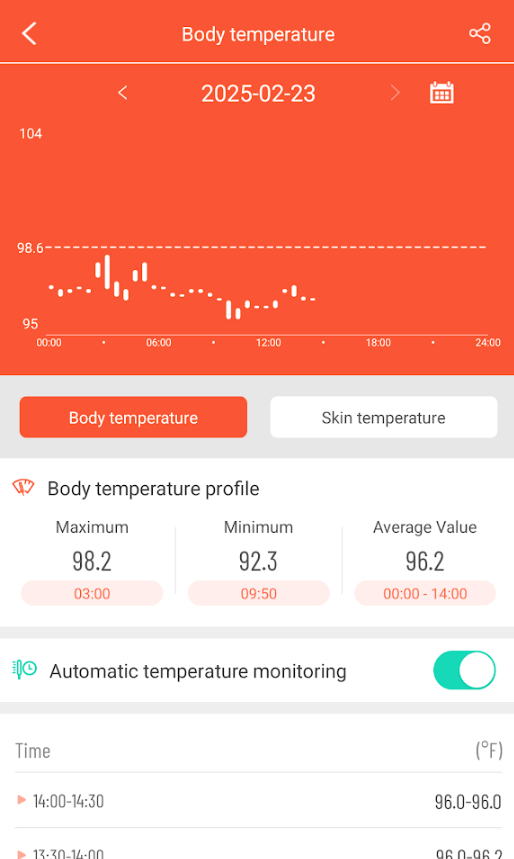
As is common with ME/CFS people in general, I continue to have sub-normal temperatures. I notice that there is often a temperature spike while I sleep.
As often happens with ME/CFS, there can be low blood pressure as well as POTS.
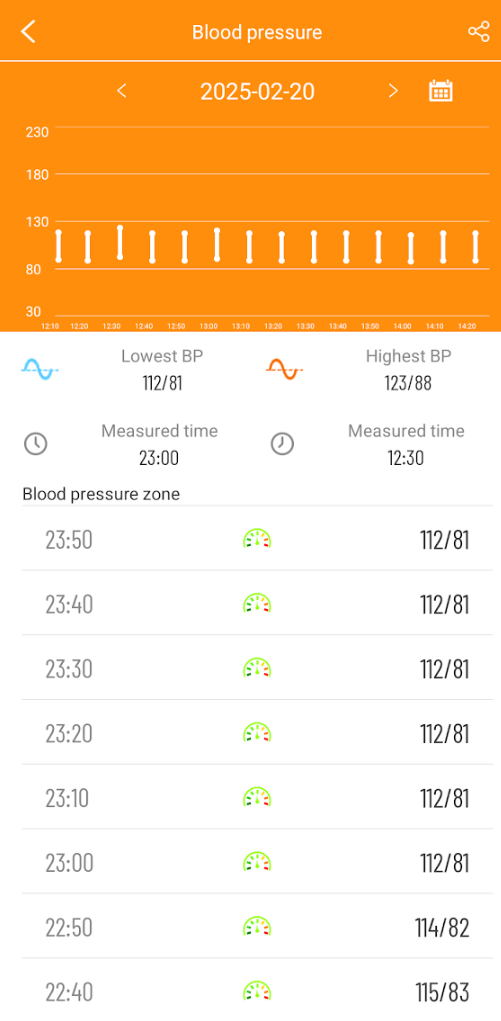
My variant of ME/CFS had coagulation being a factor. This can result in hypoxia (low oxygen levels) causing cognitive issues.

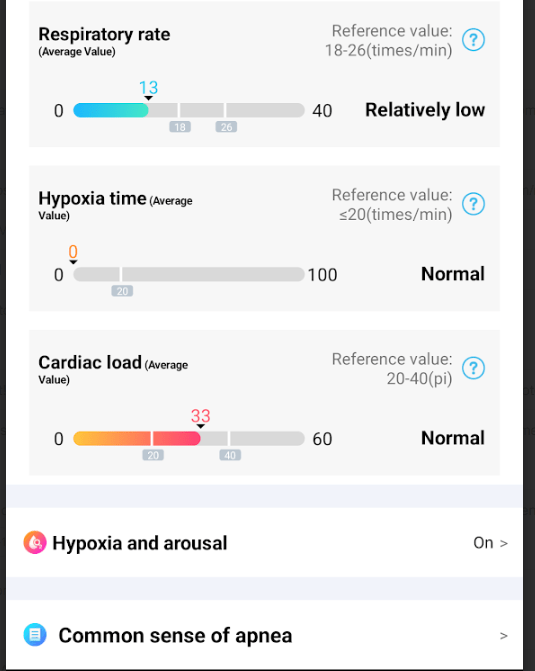

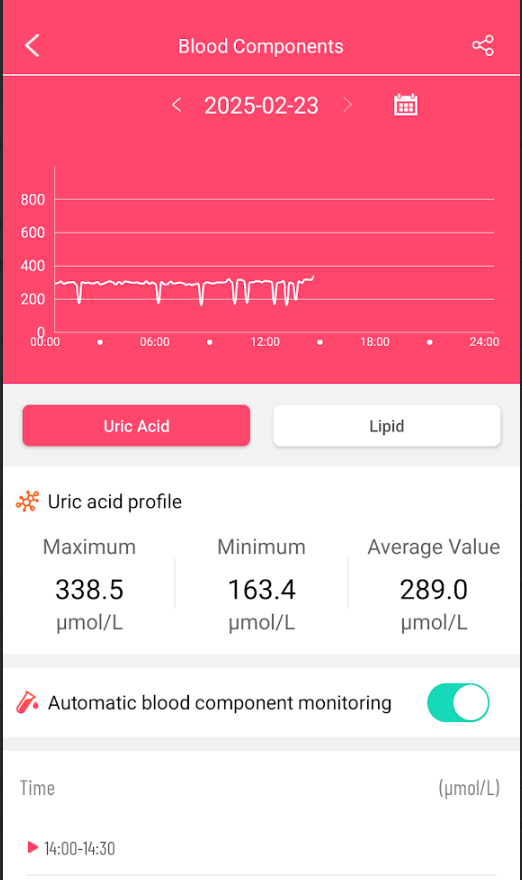


Heart rate variability (HRV) is a measure of how much time varies between heartbeats. It’s a physiological marker of heart health and the autonomic nervous system (ANS). HRV can be used to assess physical and mental health, and to help diagnose diseases.
How HRV is measured
What HRV indicates

Heart Issues are frequently seen with ME/CFS. See The Heart and Blood of the CFS Patient



For example, I need to drink more water, etc.
For less than $30, you get a lot of good information to act upon.